Inspiration
As a final-year pharmacy student and an Ml engineer, l've spent the last few years learning how drugs are discovered, developed, and brought to market. But one thing kept bothering me; what happens to the brilliant ideas that never get funded? In class and journals, I kept coming across research from independent scientists, some of whom had identified novel compounds, built models, or even completed small-scale lab tests. And yet, because they weren't employed by a major pharma company or connected to institutional grants, their work stayed stuck in limbo. One story that really hit me was about Dr. Victoria Hale, the founder of OneWorld Health, who left her job at Genentech to develop low-cost drugs for neglected diseases. She had the expertise, the compound leads, and the vision-but she had to build a non-profit drug company just to get funding. The barriers were enormous. That's when I realized: R&D doesn't lack ideas. It lacks access. What if we could change that?
What it does
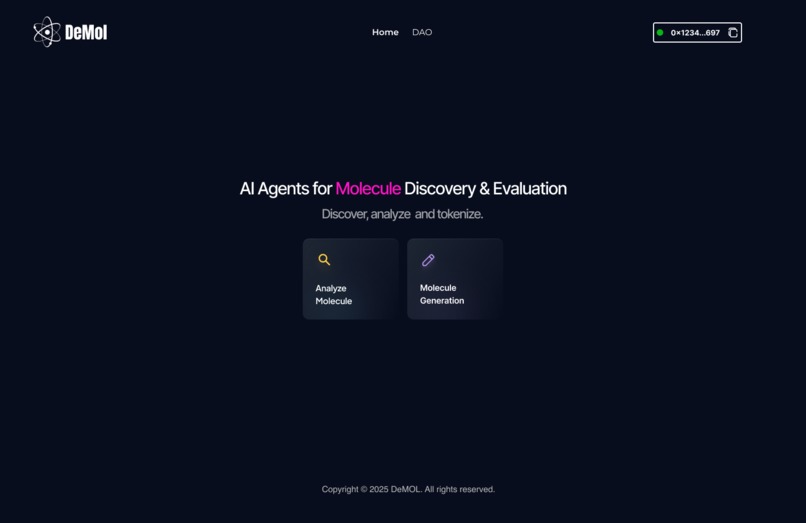
Our platform brings together AI agents, blockchain infrastructure, and DAO governance to transform how drug research happens openly, collaboratively, and transparently.
Analyze Molecules Run predictive models to evaluate activity, toxicity, drug-likeness, and synthesizability, fully verifiable and automated.
Generate Molecules Use AI agents to create novel drug-like scaffolds and explore chemical space with speed and intelligence.
Tokenize Promising Compounds Molecules that meet eligibility thresholds are minted as on-chain IP, complete with metadata and scientific scoring.
Propose & Vote The DAO reviews tokenized candidates and votes on which ones to fund for real-world research and development.
Share the Upside Researchers and DAO participants earn a stake in the success of funded compounds, with transparent royalties and governance rights.
How we built it
DeMol combines cutting-edge machine learning, decentralized infrastructure, and smart governance to reshape how drug discovery is done and funded.
- Machine Learning
We implemented several specialized ML models to power molecule evaluation and design:
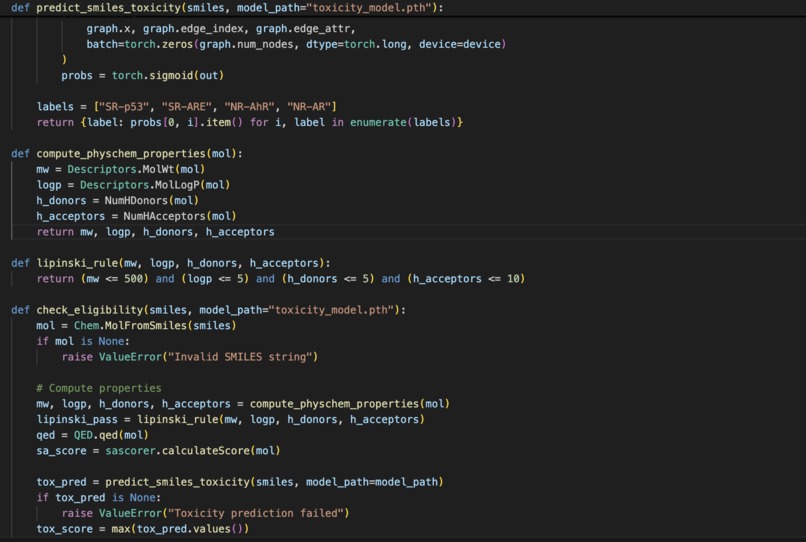
- A toxicity prediction model to flag potentially harmful candidates.
- A Lipinski + QED scoring model using AttentiveFP, implemented in PyTorch, to assess drug-likeness and developability.
- A generative model, also built in PyTorch, capable of proposing novel scaffold structures.
- A Synthetic Accessibility (SA) scoring module, based on the method by Ertl & Schuffenhauer, to assess how easily a molecule can be synthesized.
These models are orchestrated via ELiZA OS and AWS for inference, with scoring results stored and utilized in downstream tokenization logic.
- Tokenization & Blockchain
- Smart contracts (ERC-721 for molecules, ERC-20 for governance/funding) were deployed on Avalanche C-Chain and Ethereum Sepolia.
- Molecules that meet eligibility thresholds are automatically tokenized as IP NFTs and stored with metadata on AWS S3.
- Chainlink Automation handle score fetching, minting triggers, and DAO workflow logic.
- Chainlink CCIP enables future interoperability across chains (e.g., Optimism, Arbitrum).
Tokenized IP is DAO-governed and optionally yield-bearing, with infrastructure in place for royalty streaming (e.g., via Superfluid).
DAO & Proposal System
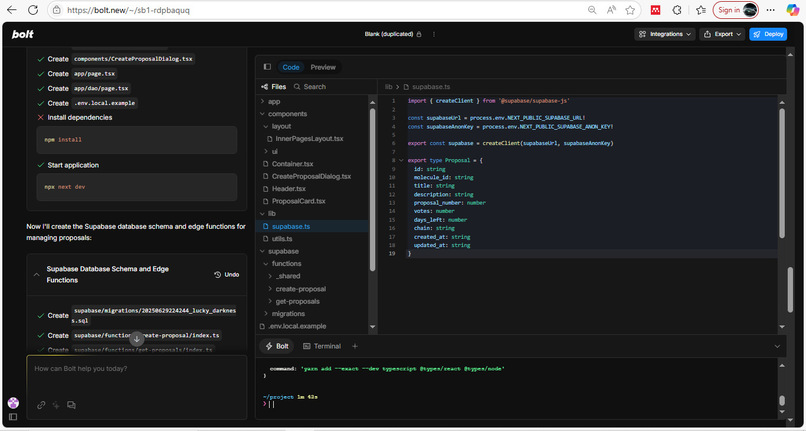
Supabase is used to store and retrieve proposal data, enabling fast, reliable off-chain access for the UI and governance history.
A DAO proposal and voting system lets the community fund real-world R&D of high-scoring molecules.
DAO metadata and proposal content are stored in Supabase, enabling a fast and reliable off-chain record for the UI and governance history.
The DAO can vote to release funds to researchers or partner labs when milestones are met.
Frontend & User Interface
The UI was initially designed in Figma, with a strong focus on clarity, accessibility, and scientific credibility.
We built the interface using Bolt.new’s AI-powered development platform, which accelerated the frontend workflow by generating high-quality, production-ready React components.
The frontend stack includes Next.js, TailwindCSS, and Framer Motion, delivering a smooth and responsive user experience.
Users can submit molecules, view ML scoring results, participate in DAO governance, and interact with tokenized IP, all within a unified, elegant dApp interface.
Wallet connections and on-chain actions are handled seamlessly using wagmi and ethers.js, ensuring secure and efficient blockchain interaction.
The dApp is deployed on Netlify, providing fast, scalable, and reliable hosting for global users.
AI Agent Orchestration (ELiZA OS)
Key ML models are deployed as ELiZA agents, allowing verifiable, modular AI scoring pipelines.
ELiZA ensures that molecule scoring and downstream decisions (like minting or proposal creation) are transparent, traceable, and cryptographically verifiable.
Agent outputs are directly tied to blockchain actions, closing the loop between computation and coordination.
Challenges we ran into
- ELiZA Inference Mapping: Customizing agent actions to be recognized and executed correctly within ELiZA’s framework.
- *Lightweight Deployment: * Running ELiZA agents on small servers (e.g., AWS micro instances) caused performance bottlenecks.
- *Bolt.new Integration: * Adapting AI-generated components to match Figma designs and support complex Web3 logic.
- *Toxicity Model Balance: * Managing performance across multiple toxicity endpoints with limited data.
Accomplishments that we're proud of
- Built a functional AI pipeline that generates and analyzes drug-like molecules using scaffold-based design.
- Integrated tokenization logic to mint eligible compounds as on-chain molecular IP with full metadata.
- Connected predictive models as modular ELiZA agents, ready for verifiable execution and coordination.
- Launched on Avalanche, ensuring fast, low-cost transactions for molecule minting and DAO actions.
- Implemented Chainlink CCIP & Automation, enabling cross-chain compatibility and real-world data triggers.
- Designed a clean and intuitive UI that makes drug discovery accessible to researchers, scientists, and the Web3 community.
What we learned
- Building AI Agents with ELiZA OS: Learned how to wrap ML models into modular, verifiable agents and connect them to on-chain logic.
- Debugging Chainlink Automation Locally: Gained experience simulating and testing Chainlink Functions and Automation without full mainnet deployment.
- Cross-Chain Interactions via CCIP: Understood how to securely pass messages and tokens between Avalanche and Ethereum using Chainlink CCIP.
- Using Bolt.new for UI Development: Learned how to effectively guide an AI-based UI generator, then customize it for dApp requirements.
What's next for DeMol
- Structure-based drug design
- Target-specific molecule generation
- Multi-omics data interpretation
- Synthesis cost prediction and route planning
- Partnerships with independent researchers, universities, and biohackers to onboard promising molecules from underfunded labs.
- Educational content and tools to help new contributors understand the system and get involved in molecule generation, analysis, and voting.
- Launch of researcher reputation scores to spotlight credible contributors.
- Community-curated review boards for more transparent and expert-led proposal evaluation.
- Cross-chain functionality powered by Chainlink CCIP, already live on Avalanche and Ethereum Sepolia, laying the groundwork for seamless interactions with major ecosystems like Optimism, Arbitrum, and beyond.
Built With
- avalanchec-chain
- bolt.new
- chainlinkautomation
- chainlinkccip
- docker
- elizaos
- erc1155
- etherscan
- foundry
- framermotion
- netlify
- nextjs
- openrouter
- openzeppelin
- python
- pythonflask
- rdkit
- react
- solidity
- supabase
- tailwindcss
- torch
- typescript

Log in or sign up for Devpost to join the conversation.