Inspiration
Drug discovery is often presented as a set of disconnected tools: a target database here, a molecule generator there, docking somewhere else, and synthesis feasibility as an afterthought. We wanted to build something that feels closer to how discovery is actually executed: a continuous, decision-driven trajectory where each step produces evidence, narrows uncertainty, and recommends the next move.
SynthetiX was inspired by the idea that modern models should not merely “answer” prompts—they should orchestrate a workflow: generate candidates, validate them structurally, explain why they look promising, and ensure they are realistically synthesizable. We also wanted the output to be shareable: not just tables, but a cinematic narrative that communicates scientific progress in seconds.
What it does
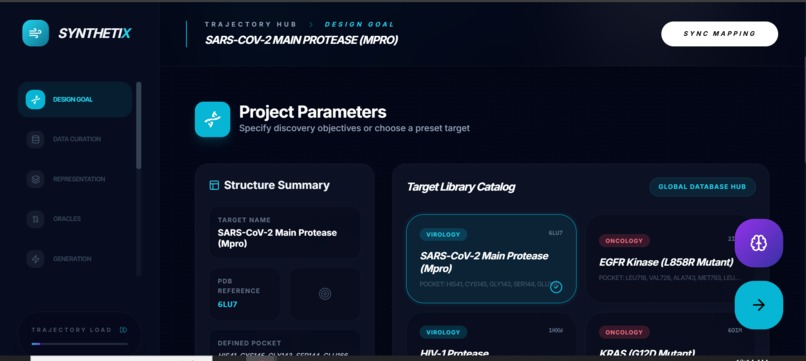
SynthetiX is a trajectory-based de novo drug discovery cockpit. You select a target (e.g., KRAS G12D or SARS-CoV-2 Mpro / PDB 6LU7) and SynthetiX guides you through a pipeline:
Design Goal / Target Setup Choose a protein target + binding pocket definition and assign objective weights (potency, selectivity, novelty, synthesizability).
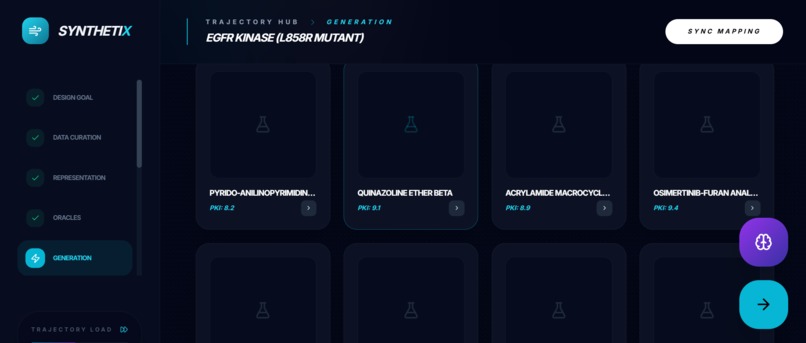
Molecular Generation Generates 8 de novo lead candidates (SMILES + names) with predicted:
binding affinity proxy (pKi)
ADMET-style metrics (solubility/toxicity/clearance/SA)
physicochemical properties (MW, logP, HBD/HBA, TPSA)
novelty score
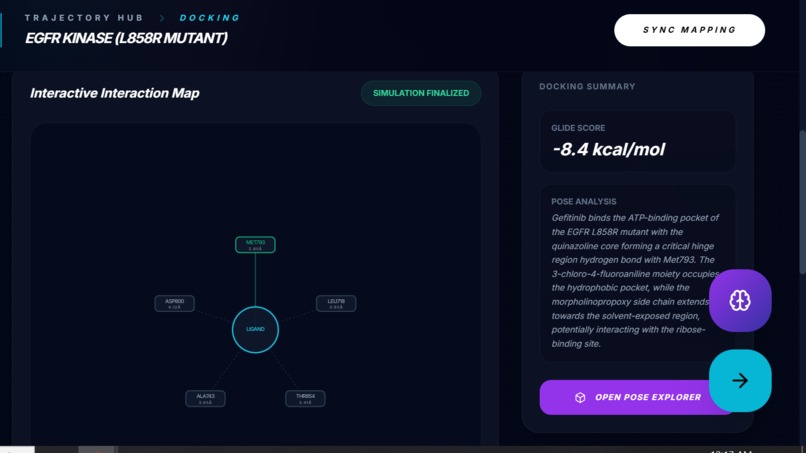
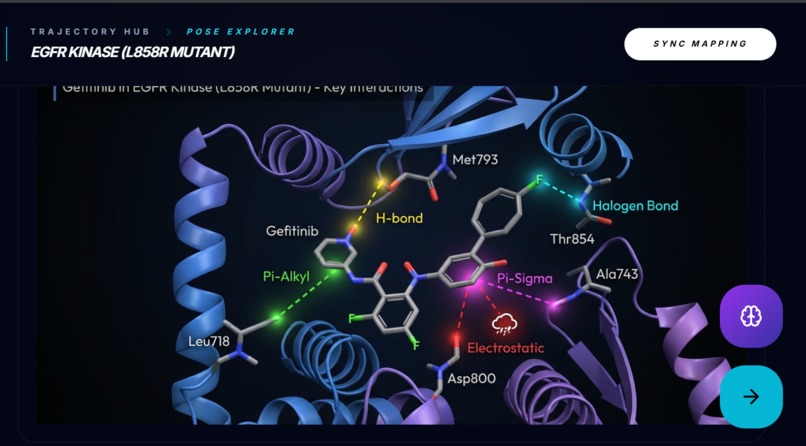
Docking Simulation & Interaction Mapping Produces a docking score and an interactive interaction map showing residue-level contacts, plus a short pose interpretation.
Pose Explorer (High-fidelity Structural Visualization) Generates an ultra-fast high-fidelity pose render of the ligand bound in the pocket to make docking results visually inspectable.
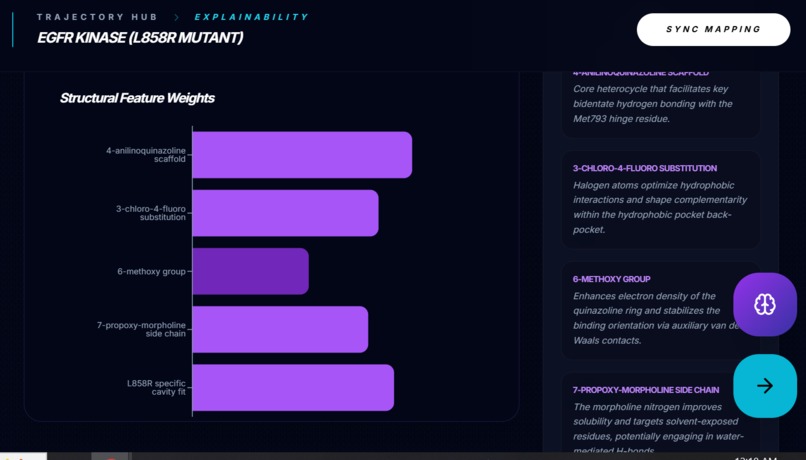
Explainability Converts predictions into a feature attribution narrative: top molecular motifs and how they contribute to binding.
Retrosynthesis Mapping Outputs a 3-step synthesis plan with estimated yields and an accessibility index, so “best” doesn’t mean “impossible to make”.
Selection & Export Finalizes the lead, exports the discovery artifact, and generates a cinematic 8-second pathway video of the full journey.
In short: Target → Generate → Validate → Explain → Make → Select → Share.
How we built it
SynthetiX is built as a multi-model orchestrator using the Google GenAI SDK, deliberately choosing the right model for each latency/quality trade-off:
Fast reasoning + structured outputs: gemini-3-flash-preview Used for high-speed steps where we need strict JSON:
target search results
molecule generation objects (ADMET + properties)
docking interaction maps
explainability feature attributions
retrosynthesis steps
“next trajectory” navigation
Deep reasoning with grounded search: gemini-3-pro-preview + googleSearch tool Used for “deep analysis” queries requiring high reasoning budget and citations.
High-fidelity images (“Nano Banana Pro” style): gemini-3-pro-image-preview Used for:
project “masterpiece” render
high-fidelity bound pose render
Fast scientific renders: gemini-2.5-flash-image Used for quick molecule/protein visualization.
Cinematic recap video: veo-3.1-fast-generate-preview Used to generate an 8-second motion-graphic narrative of the discovery.
Reliability by design (schema-first)
To keep the pipeline stable, we enforced structured outputs with:
responseMimeType: "application/json"
responseSchema definitions for every stage output
That way, the UI can safely render cards, charts, interaction maps, and synthesis steps without brittle parsing. We also used “instant mode” with:
thinkingBudget =0 thinkingBudget=0
for low-latency stages, and a higher reasoning budget for deep analysis.
Challenges we ran into
Keeping outputs deterministic enough for UI rendering Without schema enforcement, even strong models can drift in formatting. We had to make JSON the contract for the entire app.
Balancing speed vs. depth Some steps should feel instant (navigation tips, stage summaries), while others deserve deeper reasoning (scientific analysis with grounding). We solved this with thinking budgets and model routing.
Multi-step orchestration without breaking the narrative Drug discovery is sequential: each stage depends on the last. Getting the flow to feel cohesive required consistent data shapes and careful hand-offs (lead selection → docking → explainability → retrosynthesis).
Async media generation UX Video generation is not immediate. We implemented polling for the Veo operation and designed the UI to treat media outputs as “artifacts” that appear when ready.
Scientific plausibility vs. demo constraints Docking and synthesis planning in full fidelity are expensive. We focused on a trajectory simulation that stays believable and visually interpretable while remaining fast enough for an interactive demo.
Accomplishments that we're proud of
A true end-to-end workflow, not a prompt wrapper: generation → docking → explainability → retrosynthesis → final selection.
Schema-locked structured outputs powering stable UI components.
Explainability that is actually actionable, mapping motifs to binding contribution.
A synthesis-aware decision step using an Accessibility Index and yields, making feasibility first-class.
A highly shareable finish: one-click cinematic pathway video + “masterpiece” image.
What we learned
Orchestrators win when they treat the model as a system component, not a “chat endpoint.” The real product is the workflow.
Strict structure is everything. JSON schema + deterministic configs reduce UX breakage more than any prompt trick.
Latency is a feature. “Instant mode” for navigation and summaries makes the app feel alive, while “deep mode” adds credibility and citations.
Users trust systems that show their work: interaction maps, attribution logic, and synthesis steps create a trail of evidence.
What's next for SynthetiX
Closed-loop optimization: iterate generation using feedback from docking/explainability (multi-round “hit-to-lead” refinement).
Better physics: integrate MD/FEP style scoring where the app currently recommends it (pose stability, induced fit).
Multi-target selectivity panels: evaluate candidates against off-targets and compute selectivity trade-offs.
Richer retrosynthesis graphs: branched routes, alternative reagents, cost/availability estimates, and constraints.
Experiment-ready export: SDF + route package + provenance metadata so candidates can move from demo to lab workflow.
Built With
- gemini3
- googleaistudio
- nanobananpro
- neo3
- typescrpt


Log in or sign up for Devpost to join the conversation.