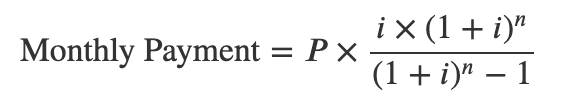MIMS was introduced in the NHANES Study
A lot the accelerometry data from ActiGraph devices is summarized
using ActiLife software into ActiGraph Activity Counts (AC). The NHANES
(National Health and Nutrition Examination Survey) used ActiGraph devices, collecting data on over 14000 people in a nationally representative survey. The NHANES introduced MIMS, a Monitor-Independent
Movement Summary for activity data (John et al.
2019) instead of AC. The NHANES study released these summaries in
large (over 7Gb) SAS
XPORT files (XPTs), found in the Examination data. As this was a
novel measure, Karas et al. (2022)
compared MIMS and other common summary measures, which was possible
since MIMSunit package in R implements this method. In
2022, NHANES released their raw 80Hz accelerometer data, which was large
(over 1Tb compressed). We can use this raw data to compute MIMS using
the method described in John et al. (2019)
and compare it to the released data. **In short, we found that the
default MIMSunit::mims_unit
does not calculate the same result as released by NHANES, you must use
the MIMSunit::custom_mims_unit
function with allow_truncation = FALSE to get almost exact
results.
Here we’re showing this fact with one participant. Note, this code
may take some time to download the data, but we will walk through each
step using a reprex approach
and the full code is available at https://github.com/muschellij2/HopStat/blob/gh-pages/NHANES_Activity_using_MIMS_(Monitor-Independent_Movement_Summary)/NHANES_Activity_using_MIMS_(Monitor-Independent_Movement_Summary).Rmd.
Package Loading
We’re going to load packages required for this analysis:
library(MIMSunit)
library(curl)
library(haven)
library(curl)
library(dplyr)
library(readr)
options(digits.secs = 3)Set up
The code below can do all of this in a temporary directory. I used a
hard-coded directory since some of these files are big and take a while
to download/convert/run. If you want to do this in a temporary way, you
can run:
Otherwise, set tdir to be a directory on your machine
for the data to go.
Here we pick a random ID. This ID is technically in the NHANES
National Youth Fitness Survey (NNYFS), but the same processing and
analysis was done on this data compared to NHANES. An ID from this study
was chosen since it has only around 1700 participants, and so the XPT is
smaller to download and load into R.
id = "71917"Downloading Raw Data
The raw data are given in series of zipped tarballs, one for each
person, and the code below will download the tarball for the ID above.
The function below is a general function to download any of the tarballs
from the CDC FTP site. The
version argument is the folder on the FTP site
(pax_y, pax_g or pax_h), and
id is the ID for the person, also referred to the
SEQN in NHANES documentation. The exdir
argument is where to put the downloaded files, and ... are
additional arguments passed to curl::curl_download
such as quiet = TRUE/FALSE.
# Download tarball
download_80hz = function(id, version, exdir = tempdir(), ...) {
files = id
tarball_ending = grepl("[.]tar[.]bz2", id)
files[!tarball_ending] = paste0(files[!tarball_ending], ".tar.bz2")
urls = paste0("https://ftp.cdc.gov/pub/", version, "/", files)
outfiles = sapply(urls, function(x) {
destfile = file.path(exdir, basename(x))
if (!file.exists(destfile)) {
curl::curl_download(x, destfile = destfile, ...)
}
destfile
})
outfiles
}Here we download the tarball for the data (about 130Mb).
raw = download_80hz(id, "pax_y", quiet = FALSE, exdir = tdir)Reading in the Raw data
Here we read the data and write it out as a CSV, so we do not have to
re-run the extraction from the tarball, which can take a little bit of
time, in future execution. The tarball has a series of CSV files, one
for each hour, and one log file. We exclude the log file and read in the
CSVs using readr.
file_data = file.path(tdir, paste0(id, ".csv"))
if (!file.exists(file_data)) {
files = untar(raw, verbose = TRUE, exdir = tdir, list = TRUE)
files = file.path(tdir, files)
if (!all(file.exists(files))) {
untar(raw, verbose = TRUE, exdir = tdir)
}
log_file = files[grepl("_log", files, ignore.case = TRUE)]
#' Getting only the data files
csv_files = files[!grepl("_log", files, ignore.case = TRUE)]
df = readr::read_csv(
csv_files,
col_types = readr::cols(
HEADER_TIMESTAMP = readr::col_datetime(),
X = readr::col_double(),
Y = readr::col_double(),
Z = readr::col_double()
)
)
readr::write_csv(df, file_data)
} else {
df = readr::read_csv(
file_data,
col_types = readr::cols(
HEADER_TIMESTAMP = readr::col_datetime(),
X = readr::col_double(),
Y = readr::col_double(),
Z = readr::col_double()
),
progress = FALSE,
show_col_types = FALSE
)
}Raw Data
Here we can print out the data to see what it looks like. We have it
in a tibble so it will not print all records and the
options(digits.secs = 3) set above will show the
milliseconds for the time of the data:
df# A tibble: 55,343,278 × 4
HEADER_TIMESTAMP X Y Z
<dttm> <dbl> <dbl> <dbl>
1 2000-01-07 13:25:00.000 -0.774 0.009 0.604
2 2000-01-07 13:25:00.013 -0.739 0.018 0.569
3 2000-01-07 13:25:00.024 -0.704 0.023 0.522
4 2000-01-07 13:25:00.036 -0.748 0.023 0.449
5 2000-01-07 13:25:00.049 -0.801 0.029 0.384
6 2000-01-07 13:25:00.062 -0.83 0.032 0.34
7 2000-01-07 13:25:00.075 -0.845 0.038 0.346
8 2000-01-07 13:25:00.088 -0.874 0.029 0.34
9 2000-01-07 13:25:00.100 -0.891 0.026 0.314
10 2000-01-07 13:25:00.113 -0.918 0.018 0.27
# ℹ 55,343,268 more rowsWe see the data is made up of time (HEADER_TIMESTAMP),
and the accelerometry values for the 3 axes, measures in g
(gravitational units, \(g =
9.81m/s^2\)). The data is collected at 80Hz, so there are 80 rows
per second.
Note: the data column name is
HEADER_TIMESTAMP, but the MIMSunit package assumes the time
column is called HEADER_TIME_STAMP (note the extra
underscore).
With the lubridate package, we can floor the time to the
second level, count the number of rows with that time value and see that
the majority are 80, showing 80Hz data, with some that do
not have a full second covered.
df %>%
dplyr::mutate(time = lubridate::floor_date(HEADER_TIMESTAMP, unit = "second")) %>%
dplyr::count(time) %>%
dplyr::count(n, name = "n_samples")# A tibble: 2 × 2
n n_samples
<int> <int>
1 78 1
2 80 691790Calculating MIMS
From this df data, we can calculate MIMS using the
MIMSunit package. This package has a function mims_unit
which is the default way to calculate MIMS, and then use custom_mims_unit
function which allows more options. We will use both of these functions
to calculate MIMS and compare them to the released MIMS data from
NHANES. Note again the renaming of columns. This process interpolates
the data to 100Hz, runs an extrapolation procedure depending on the
dynamic range of the device (\(\pm
6g\)) for samples that reach the boundary, then applies a
4th-order Butterworth filter with cutoffs at 0.2 to 5Hz (limiting to
“typical” human activity) and then integrates the area under the curve
for each axis (MIMS per axis). The sum of these MIMS per axis gives a
minute level MIMS. This is done for each epoch, which in NHANES is 1
minute. See John et al. (2019) for more
details.
file_mims = file.path(tdir, paste0(id, "_MIMS.csv"))
if (!file.exists(file_mims)) {
# MIMSunit requires specific naming
run_df = df %>%
dplyr::rename(HEADER_TIME_STAMP = HEADER_TIMESTAMP)
mims = MIMSunit::mims_unit(df, epoch = "1 min", dynamic_range = c(-6L, 6L),
output_mims_per_axis = TRUE)
# for printing
mims = as_tibble(mims)
readr::write_csv(mims, file_mims)
} else {
mims = readr::read_csv(
file_mims,
col_types = readr::cols(
HEADER_TIME_STAMP = readr::col_datetime(),
.default = readr::col_double()
),
progress = FALSE,
show_col_types = FALSE
)
}
mims# A tibble: 11,530 × 5
HEADER_TIME_STAMP MIMS_UNIT MIMS_UNIT_X MIMS_UNIT_Y MIMS_UNIT_Z
<dttm> <dbl> <dbl> <dbl> <dbl>
1 2000-01-07 13:25:00.000 27.3 8.41 11.7 7.13
2 2000-01-07 13:26:00.000 41.2 14.9 14.8 11.5
3 2000-01-07 13:27:00.000 30.6 10.7 10.2 9.69
4 2000-01-07 13:28:00.000 7.83 2.73 2.84 2.26
5 2000-01-07 13:29:00.000 35.3 13.4 9.61 12.3
6 2000-01-07 13:30:00.000 6.81 2.08 1.71 3.02
7 2000-01-07 13:31:00.000 27.0 8.09 9.33 9.59
8 2000-01-07 13:32:00.000 30.2 9.12 11.8 9.31
9 2000-01-07 13:33:00.000 9.71 2.60 3.61 3.50
10 2000-01-07 13:34:00.000 16.0 4.88 6.43 4.72
# ℹ 11,520 more rowsCalculating MIMS without truncation
Here we will run the same procedure, but with
allow_truncation = FALSE, which, according to the
documentation
. If it is TRUE, the algorithm will truncate very small MIMS-unit values to zero.
The threshold is based on
1e-04 * parse_epoch_string(epoch, sr) from this
code.
As the data is interpolated to 100Hz (so sr = 100), and
the epoch is 1 min, this threshold is:
1e-04 * MIMSunit::parse_epoch_string("1 min", sr = 100)[1] 0.6So that any minute-level MIMS value below 0.6 is
truncated to 0 (when using mims_unit default).
This is a very small value, but it does have an effect on the results,
as we will see below.
file_mims_notrunc = file.path(tdir, paste0(id, "_MIMS_notrunc.csv"))
if (!file.exists(file_mims_notrunc)) {
# MIMSunit requires specific naming
run_df = df %>%
dplyr::rename(HEADER_TIME_STAMP = HEADER_TIMESTAMP)
# no allowing truncation
mims_notrunc = MIMSunit::custom_mims_unit(
run_df, epoch = "1 min",
dynamic_range = c(-6L, 6L),
output_mims_per_axis = TRUE,
allow_truncation = FALSE)
# for printing
mims_notrunc = as_tibble(mims_notrunc)
readr::write_csv(mims_notrunc, file_mims_notrunc)
} else {
mims_notrunc = readr::read_csv(
file_mims_notrunc,
col_types = readr::cols(
HEADER_TIME_STAMP = readr::col_datetime(),
.default = readr::col_double()
),
progress = FALSE,
show_col_types = FALSE
)
}mims_notrunc# A tibble: 11,530 × 5
HEADER_TIME_STAMP MIMS_UNIT MIMS_UNIT_X MIMS_UNIT_Y MIMS_UNIT_Z
<dttm> <dbl> <dbl> <dbl> <dbl>
1 2000-01-07 13:25:00.000 27.3 8.41 11.7 7.13
2 2000-01-07 13:26:00.000 41.2 14.9 14.8 11.5
3 2000-01-07 13:27:00.000 30.6 10.7 10.2 9.69
4 2000-01-07 13:28:00.000 7.83 2.73 2.84 2.26
5 2000-01-07 13:29:00.000 35.3 13.4 9.61 12.3
6 2000-01-07 13:30:00.000 6.81 2.08 1.71 3.02
7 2000-01-07 13:31:00.000 27.0 8.09 9.33 9.59
8 2000-01-07 13:32:00.000 30.2 9.12 11.8 9.31
9 2000-01-07 13:33:00.000 9.71 2.60 3.61 3.50
10 2000-01-07 13:34:00.000 16.0 4.88 6.43 4.72
# ℹ 11,520 more rowsWe see that the data is highly correlated regardless of options
used:
cor(mims$MIMS_UNIT, mims_notrunc$MIMS_UNIT)[1] 0.9998119But we do see some differences, with a maximum absolute difference of
> 1, which for MIMS is significant:
hist(abs(mims$MIMS_UNIT - mims_notrunc$MIMS_UNIT))MIMS released by NHANES
Here we download the MIMS data released from NHANES from the XPT
(1.6Gb – large).
file_mims_xpt = file.path(tdir, paste0(id, "_MIMS_XPT.csv"))
if (!file.exists(file_mims_xpt)) {
url = "https://wwwn.cdc.gov/Nchs/Data/Nnyfs/Public/2012/DataFiles/Y_PAXMIN.xpt"
destfile = file.path(tdir, basename(url))
if (!file.exists(destfile)) {
curl::curl_download(url, destfile = destfile, quiet = FALSE)
}
#' We can read in the XPT using `haven` and then subset the ID we need. We chose the first ID
pax = haven::read_xpt(destfile, n_max = 50000)
stopifnot(id %in% pax$SEQN)
#' Keep only the ids we need from above
paxmims = pax %>%
filter(SEQN %in% id) %>%
select(SEQN, PAXDAYM, PAXDAYWM, PAXSSNMP, PAXTSM, PAXAISMM, PAXMTSM, PAXMXM, PAXMYM, PAXMZM)
readr::write_csv(paxmims, file_mims_xpt)
} else {
paxmims = readr::read_csv(
file_mims_xpt,
col_types = readr::cols(
SEQN = col_character(),
PAXDAYM = col_character(),
PAXDAYWM = col_character(),
.default = readr::col_double()
),
progress = FALSE,
show_col_types = FALSE
)
}This data doesn’t have any times in it so we can check to make sure
they have the same rows and simply bind them. This would likely need
more checking for other data, but here we see these agree. Aside: A
mapping from this to the date/time data would be helpful.
head(paxmims)# A tibble: 6 × 10
SEQN PAXDAYM PAXDAYWM PAXSSNMP PAXTSM PAXAISMM PAXMTSM PAXMXM PAXMYM PAXMZM
<chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 71917 1 6 0 60 0 27.3 8.41 11.7 7.13
2 71917 1 6 4800 60 0 41.2 14.9 14.8 11.5
3 71917 1 6 9600 60 0 30.6 10.7 10.2 9.69
4 71917 1 6 14400 60 80 7.83 2.73 2.84 2.26
5 71917 1 6 19200 60 0 35.3 13.4 9.61 12.3
6 71917 1 6 24000 60 640 6.81 2.08 1.71 3.02Note, the PAXMTSM is the MIMS value released by NHANES,
the other columns are MIMS per axis (PAXMXM – X-axis MIMS),
The PAXSSNMP is the time in samples from the start of the
measurement, so 0 is the first minute, 4800 is
the second minute (80Hz * 60seconds), and PAXDAYM is the
day of measurement, so 1 is the first day, 2
is the second day, etc.
We can see that this participant was observed for a little over 8
days:
max(paxmims$PAXSSNMP)/80/60/60/24[1] 8.00625which we can confirm in the raw data
range(df$HEADER_TIMESTAMP)[1] "2000-01-07 13:25:00.000 UTC" "2000-01-15 13:34:50.963 UTC"diff(range(df$HEADER_TIMESTAMP), units = "days")Time difference of 8.00684 daysCompare Calculated MIMS to Released MIMS
Here we will check to make sure they have the same rows and simply
bind them. In practice, this may be off by 1 since calculated MIMS may
have a trailing value of -0.01 for minutes that are not
fully covered, but these will be dropped in the released MIMS:
stopifnot(nrow(paxmims) == nrow(mims))We can check the first few rows to see indeed they have very similar
results:
head(paxmims %>% select(PAXMTSM))# A tibble: 6 × 1
PAXMTSM
<dbl>
1 27.3
2 41.2
3 30.6
4 7.83
5 35.3
6 6.81head(mims %>% select(HEADER_TIME_STAMP, MIMS_UNIT))# A tibble: 6 × 2
HEADER_TIME_STAMP MIMS_UNIT
<dttm> <dbl>
1 2000-01-07 13:25:00.000 27.3
2 2000-01-07 13:26:00.000 41.2
3 2000-01-07 13:27:00.000 30.6
4 2000-01-07 13:28:00.000 7.83
5 2000-01-07 13:29:00.000 35.3
6 2000-01-07 13:30:00.000 6.81head(mims_notrunc %>% select(HEADER_TIME_STAMP, MIMS_UNIT))# A tibble: 6 × 2
HEADER_TIME_STAMP MIMS_UNIT
<dttm> <dbl>
1 2000-01-07 13:25:00.000 27.3
2 2000-01-07 13:26:00.000 41.2
3 2000-01-07 13:27:00.000 30.6
4 2000-01-07 13:28:00.000 7.83
5 2000-01-07 13:29:00.000 35.3
6 2000-01-07 13:30:00.000 6.81Comparing to Default (no truncation): Differences!
First we will compare the default mims_unit function
results to the released NHANES MIMS. We see that the maximum absolute
difference has some large values for MIMS, and the histogram shows a lot
of small differences, but also some large differences.
max(abs(mims$MIMS_UNIT - paxmims$PAXMTSM))[1] 1.652cor(mims$MIMS_UNIT, paxmims$PAXMTSM)[1] 0.9998118hist(abs(mims$MIMS_UNIT - paxmims$PAXMTSM))Comparing to MIMS with Truncation: Almost Identical
Next we will compare the custom_mims_unit function
results with allow_truncation = FALSE to the released
NHANES MIMS. We see that the maximum absolute difference is very small,
and the histogram shows almost all differences are very small.
max(abs(mims_notrunc$MIMS_UNIT - paxmims$PAXMTSM))[1] 0.02853675cor(mims_notrunc$MIMS_UNIT, paxmims$PAXMTSM)[1] 1hist(abs(mims_notrunc$MIMS_UNIT - paxmims$PAXMTSM))Conclusions
To get MIMS somewhat equivalent to those released by NHANES you need
to use the allow_truncation = FALSE argument in the
custom_mims_unit function. Though this will not likely
change results significantly, this is an important reproducibility note.
The authors likely did not implement truncation in their original method
and then added this later, so it makes sense why this may not be noted
in the NHANES or MIMSunit documentation. It would be
helpful if the NHANES documentation noted this truncation issue, but we
released this code to help researchers understand differences they are
seeing if they try to reproduce the released data.
References
“An Open-Source Monitor-Independent Movement Summary for
Accelerometer Data Processing.” Journal for the Measurement
of Physical Behaviour 2 (4): 268–81.
Wanigatunga, Jiawei Bai, Ciprian M Crainiceanu, and Jennifer A Schrack.
2022. “Comparison of Accelerometry-Based Measures of Physical
Activity: Retrospective Observational Data Analysis Study.”
JMIR mHealth and uHealth 10 (7): e38077.