-
-
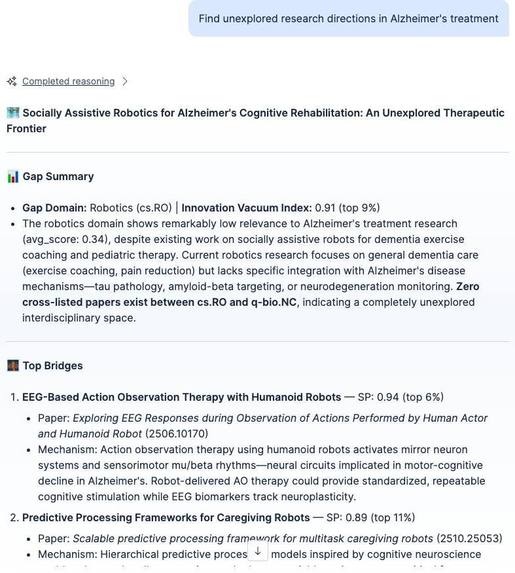
The agent's Discovery Card output: gap domain, Innovation Vacuum Index percentile, bridge candidates with Serendipity Probability scores.
-
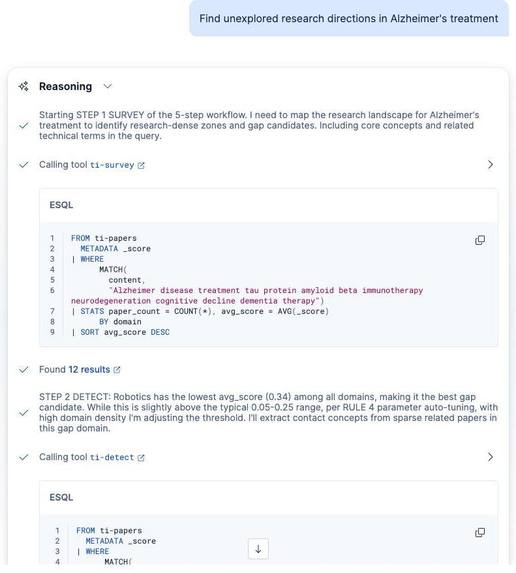
Live reasoning trace showing ES|QL tool calls (ti-survey, ti-detect), parameter auto-tuning, and step-by-step decision logging.
-
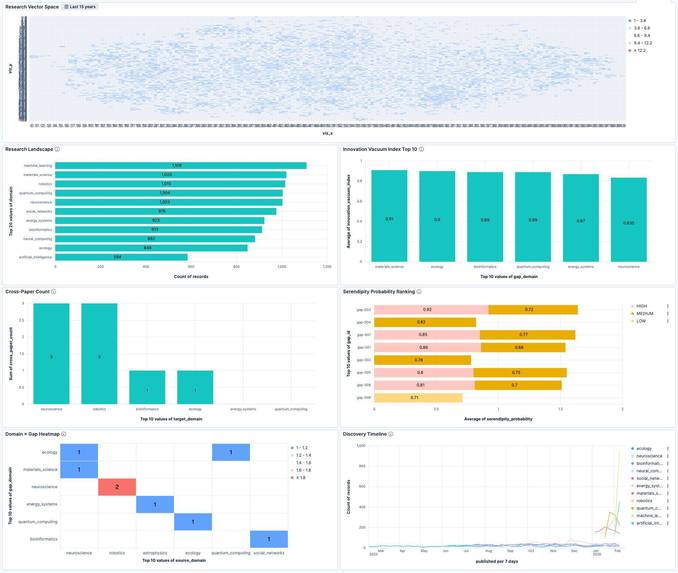
Kibana dashboard with 6 panels: Research Vector Space, IVI Top 10, Cross-Paper Count, Serendipity Ranking, Domain×Gap Heatmap, Timeline.
Inspiration
We noticed something odd about every research discovery tool available today: Semantic Scholar, Connected Papers, and Elicit all answer the same question, "What papers exist?" But some of the most important breakthroughs in science came from unexpected connections between fields. Penicillin was discovered by a bacteriologist noticing a fungal contaminant. CRISPR repurposed a bacterial immune mechanism for gene editing.
Nobody was searching for those gaps systematically. The spaces between papers, where one field has a mature solution and another field doesn't know it exists. We wanted to build a tool that finds those spaces and bridges them.
What it does
Terra Incognita is an autonomous research gap detection agent built on the Elasticsearch Agent Builder. Instead of searching for existing papers, it maps the landscape of 17,000+ scientific papers across 12 domains, identifies "meaningful voids" where research should exist but doesn't, and proposes cross-disciplinary bridges to fill them.
The agent follows a 5-step workflow (SURVEY, DETECT, BRIDGE, VALIDATE, PROPOSE), each backed by a dedicated ES|QL tool. It produces two quantitative scores:
- Innovation Vacuum Index: how meaningfully empty a gap is, displayed as percentiles like "top 2%."
- Serendipity Probability: how promising a cross-domain bridge is
Every finding is packaged as a shareable Discovery Card.
In the demo, the agent is asked about Alzheimer's treatment. It surveys 12 domains, detects a meaningful gap in materials science (IVI: top 2%), rejects a false bridge candidate ("carbon nanotube neural interfaces", keyword match only, no mechanistic relevance), and then discovers that zwitterionic anti-fouling polymers share a protein-adhesion-prevention mechanism with amyloid-beta aggregation inhibition. Zero cross-papers in our corpus. An intersection between two active fields that the agent surfaced on its own.
How we built it
The agent orchestrates 7 tools (4 ES|QL + 1 MCP + 2 platform built-in) through 9 rules defined in the system prompt. RULE/STEP numbering enforces the 5-step workflow, Self-Correction protocol, parameter auto-tuning, and Discovery Card generation.
ELSER v2 with semantic_text handles all semantic embeddings. No external model, no indexing pipeline. Just a field type declaration and 17,000+ papers become semantically searchable. This was critical for cross-domain bridge discovery: sparse vector retrieval captures meaning beyond keywords, so "protein adhesion prevention" matches "amyloid-beta aggregation" without sharing any terms.
Four parameterized ES|QL tools power the analytical backbone: per-domain relevance profiling (SURVEY), density analysis and gap identification (DETECT), cross-domain bridge search (BRIDGE), and novelty verification via cross-category count (VALIDATE).
Index Aliases enable Time-Travel backtesting: ti-papers_before_2020 for discovery, ti-papers_all for validation. The framing: "Using only data available at the time, the agent detected cross-domain signals, and subsequent papers confirmed the connection."
For automation, Cloud Scheduler triggers three daily jobs through a FastMCP server on Cloud Run: Daily Discovery (full 5-step exploration via Converse API), Gap Watch (monitor open gaps for new papers), and arXiv paper ingestion.
Challenges we ran into
The biggest challenge was making ES|QL tools write-capable. Agent Builder's ES|QL tools are read-only (SELECT only), and the Elastic Workflows execution engine had a bug in the Technical Preview where registration succeeded but execution failed. We built a custom MCP server on Cloud Run to handle all write operations, which turned out cleaner anyway.
Another challenge was avoiding false-positive bridges. Early runs would surface connections like "nano + neural" that shared keywords but had no mechanistic relevance. The Self-Correction Protocol (RULE 2) addresses this: the agent evaluates each candidate's mechanistic relevance, discards surface-level matches, and re-searches with refined concepts up to 3 times. All discard/accept decisions are recorded in the Thought Log.
The Kibana .mcp connector also doesn't forward auth headers to external MCP servers. We mitigated this with Cloud Run IAM authentication instead of app-level auth.
Accomplishments that we're proud of
The Self-Correction protocol works as intended. In the demo, the agent finds "carbon nanotube neural interfaces" as a bridge candidate, evaluates it, recognizes it's a keyword-only match with no mechanistic connection to protein aggregation, discards it, and re-searches. Then it finds zwitterionic polymers, a connection that shares no keywords with Alzheimer's research but operates on a related underlying mechanism. That rejection-then-discovery sequence is visible in the agent's output, not hidden behind the scenes.
The quantitative scoring system makes the abstract concept of "research gap" concrete and comparable. Instead of vague suggestions, researchers get percentile rankings backed by ES query results. Every score component (relevance from ELSER _score, void ratio from paper counts, density from domain statistics) traces back to a verifiable number.
We also built a fully automated daily discovery pipeline. Cloud Scheduler triggers the MCP server, which calls the Converse API to run the agent, saves results to four indices, and monitors open gaps for new publications. The system generates Discovery Cards without any human input.
What we learned
semantic_text saved us from building a separate embedding pipeline. No embedding model deployment, no vector dimension configuration, no batch indexing scripts. Just declare the field type, and ELSER handles the rest. For a hackathon project dealing with 17,000+ papers, that was a big win.
ELSER's sparse vector search turned out to be a good fit for cross-domain discovery. Dense embeddings tend to cluster by domain, but ELSER's learned sparse representations capture cross-domain semantic connections that BM25 keyword matching would miss entirely. "Anti-fouling polymer coatings" matching "amyloid-beta aggregation inhibition" is only possible because both concepts share a deeper semantic pattern (protein adhesion prevention) that ELSER captures.
Agent instruction engineering matters more than we expected. The difference between an agent that produces useful output and one that hallucinates connections came down to specific MUST/NEVER rules, step numbering, and the Self-Correction protocol. Removing platform.core.search from early iterations actually improved reliability, as the agent stopped bypassing our domain-specific tools.
What's next for Terra Incognita
The current corpus covers 12 domains and 17,000+ papers from arXiv. The immediate next step is expanding coverage: more domains, more sources (PubMed, IEEE, bioRxiv), and a larger paper corpus so the gaps the agent finds carry more weight. The 5-step workflow and Self-Correction protocol are domain-agnostic, so scaling the data doesn't require changing the agent logic.
Beyond data coverage: real-time gap alerts when a monitored gap starts filling, collaborative Discovery Cards shared across research teams, citation graph integration, and confidence calibration to track prediction accuracy over time.
The full setup scripts, MCP server, synthetic seed dataset, and arXiv collector are open-source. Other research teams can deploy the system on their own Elasticsearch clusters.
Built With
- agent-builder
- arxiv
- cloud-scheduler
- converse-api
- docker
- elasticsearch
- elser-v2
- es|ql
- fastmcp
- google-cloud-run
- index-aliases
- kibana
- python
- semantic-text
Log in or sign up for Devpost to join the conversation.