Magda Julkowska
Assistant Professor
Investigating how environmental stresses like drought and salt affect plant shape and growth, and developing innovative tools to identify crops naturally equipped to thrive in harsh conditions.

How does environmental stress shape plant architecture?
Email: mmj55@cornell.edu
Office/Lab: Room 219/212
Adjunct Assistant Professor
Plant Biology Section
School of Integrative Plant Science
Cornell University
Exogenous Hormone Treatments Reveal Species-Specific Regulation of Individual Components of Root Architecture and Salt Ion Accumulation in Cultivated and Wild Tomatoes
E Craft, M Pineros, MM Julkowska
Root remodeling mechanisms and salt tolerance trade-offs: The roles of HKT1, TMAC2, and TIP2; 2 in Arabidopsis
NO Alshareef, VJ Melino, N Saber, A De Rosa, E Rey, JY Wang, ...
PLoS genetics 21 (6), e1011713
Natural variation in salt-induced changes in root: shoot ratio reveals SR3G as a negative regulator of root suberization and salt resilience in Arabidopsis
MR Ishka, H Sussman, Y Hu, MD Alqahtani, E Craft, R Sicat, M Wang, ...
eLife 13, RP98896
Development of a mobile, high-throughput, and low-cost image-based plant growth phenotyping system
L Yu, H Sussman, O Khmelnitsky, M Rahmati Ishka, A Srinivasan, ...
Plant Physiology 196 (2), 810-829
Arabinosylation of cell wall extensin is required for the directional response to salinity in roots
Y Zou, N Gigli-Bisceglia, E van Zelm, P Kokkinopoulou, MM Julkowska, ...
The Plant Cell 36 (9), 3328-3343
We recognize that science operates within the context of wider societal, economic and political issues. We appreciate everyone’s perspectives and strive to create a safe, healthy lab environment, that allows everyone to reach their full potential, work towards their own goals, and promote the professional development of lab members at all career stages. Our lab practices include:
- Frequent mentoring sessions (for the first 3 months – once per month, afterwards every 6 months), weekly one-on-one meetings with Magda, weekly group meetings, and recognizing that clarity in instructions and expectations is kindness.
- Providing and receiving constructive feedback both ways on scientific performance, career goals, professional development and what mentoring style works for each lab member
- Normalizing and learning from failure
- Providing opportunities for science communication and participation in the scientific community through peer review, conference attendance and representation of the lab in institutional presentations and outreach activities at all career stages
- Paid undergraduate internships and overtime for non-exempt employees
- Recognizing all colleagues contain a multitude of responsibilities, obligations, and passions outside their work.
Please contact Magda for the most recent version of the lab handbook, to gain further insight into lab culture.
Research Overview
Plants adjust their development to environmental conditions. In Julkowska lab we are interested in how the environment shapes plant architecture, and what plant architectural traits contribute to overall environmental resilience. In the past, we explored Root System Architecture with simple descriptive models, like Root-Fit (Julkowska et al., 2014), as well as identified underlying genes using forward genetics (Julkowska et al., 2017). We also looked at salt stress-induced changes in root-to-shoot ratio and identified a gene of unknown function (DUF247) to play a significant role in salt stress tolerance.
The current focus of the Julkowska lab is to explore stress-induced changes in plant architecture across stress-tolerant species, including wild tomato (Solanum pimpinellifolium), cowpea (Vigna unguiculata) and tepary bean (Phaseolus acutifolius). In collaboration with Dr Chandrasekhar and Dr Lobet, we are working on new ways to describe plant architectures, by incorporating Pareto front optimality calculations (Chandrasekhar & Julkowska, 2022). Moreover, in collaboration with Dr Andrew Nelson’s lab (BTI), we are working on developing widely accessible plant phenotyping setups using Raspberry Pi and Arduino microcontrollers, and data processing pipelines based on PlantCV and R (Yu & Sussman et al., 2023).
Dr. Julkowska is the Director of BTI’s high-throughput phenotyping facility, "PhenoSight".
Please visit the BTI Plant Phenotyping Facility page to learn more about this cutting-edge resource.
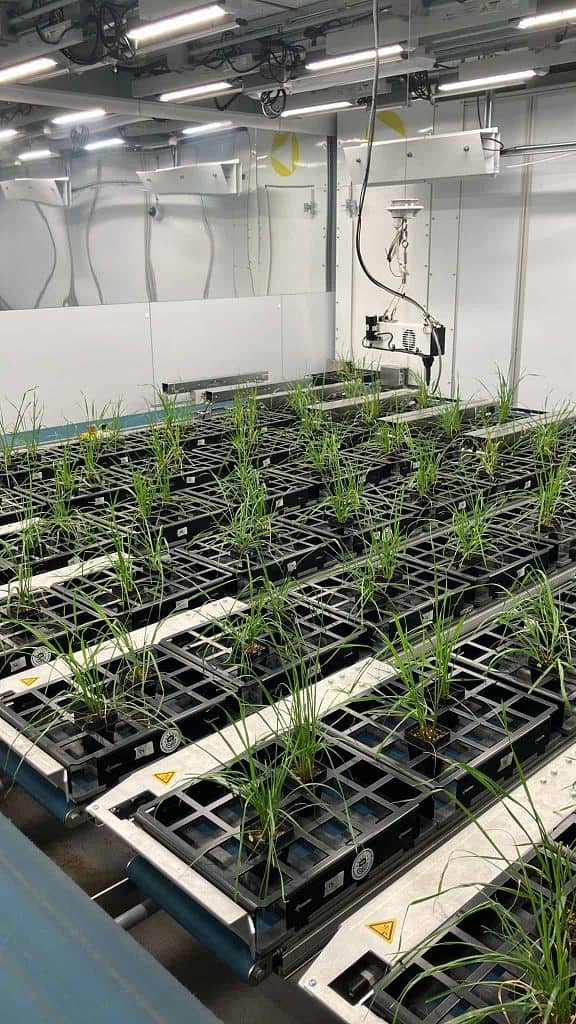
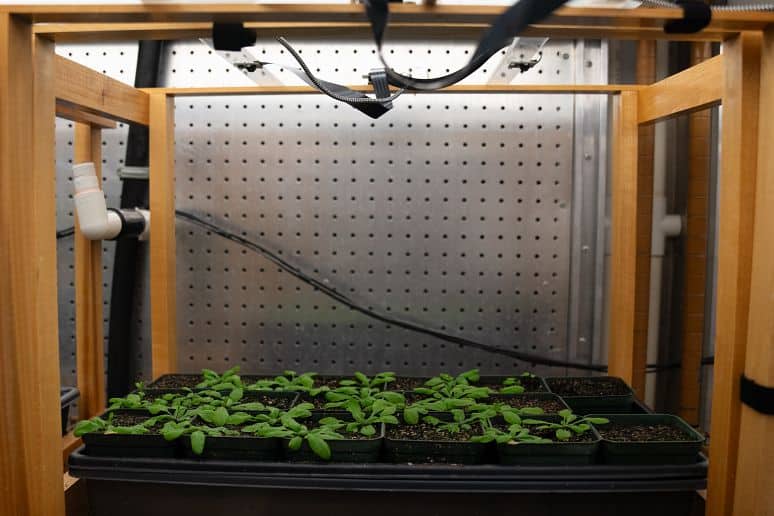
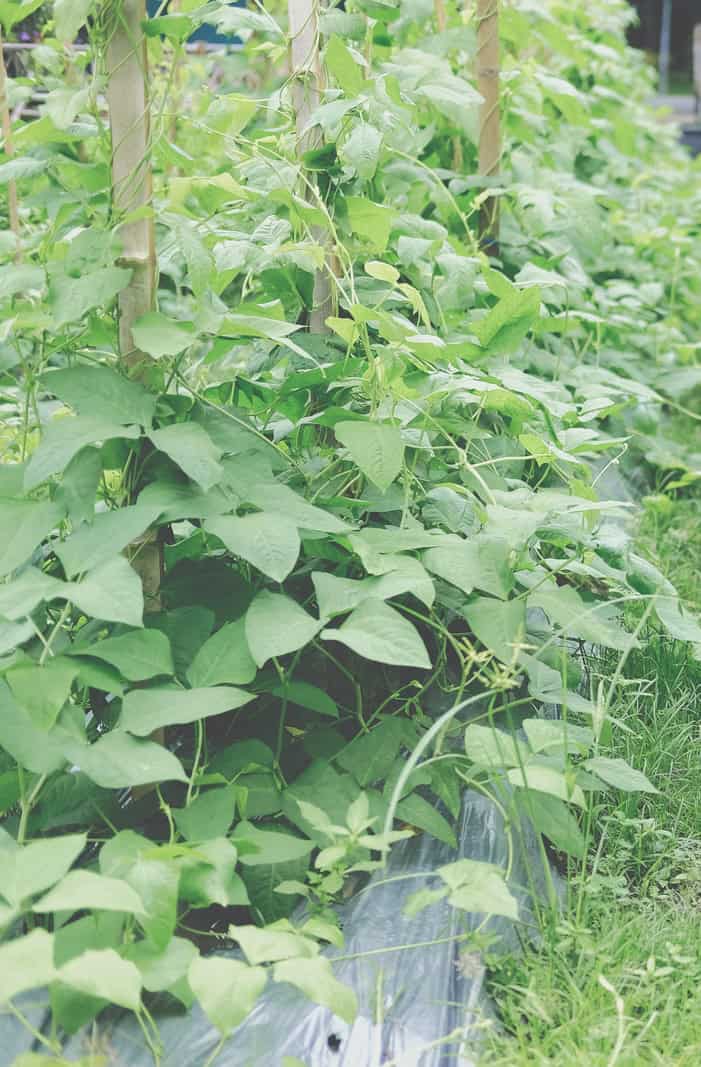




Examples of current research projects in the Julkowska Lab
 Tepary bean (Phaseolus acutifolius) is a native crop grown in arid and semi-arid regions of the Southwestern U.S., Northwestern Mexico, and Central America. While genetic and genomic resources have been developed for tepary bean breeding, we are working to improve the phenomic and genomic resources for tepary beans to accelerate breeding and discovery of genetic components associated with improved agronomic traits and environmental resilience.
Tepary bean (Phaseolus acutifolius) is a native crop grown in arid and semi-arid regions of the Southwestern U.S., Northwestern Mexico, and Central America. While genetic and genomic resources have been developed for tepary bean breeding, we are working to improve the phenomic and genomic resources for tepary beans to accelerate breeding and discovery of genetic components associated with improved agronomic traits and environmental resilience.
We isolated DNA from 400 tepary bean accessions, and genome sequencing data was used for SNP calling, in collaboration with Dr Andrew Nelson, Dr Timothy Porch (USDA-ARS) and Dr Duke Pauli (Uni. of Arizona). The entire collection was also screened for drought-induced changes in the shoot growth and architecture using the Raspberry Pi PhenoCage setup, and salt-induced changes in the root growth and architecture using the paperponics setup developed in Dr Miguel Pineros’ lab (USDA-ARS). The results of this project will be used for establishing Genome-Wide Association Study pipelines, and identifying genetic components underlying plant architecture, and stress resilience.
 Cowpea (Vigna unguiculata) is a legume crop native to West Africa, resilient to many biotic and abiotic factors. Hayley Sussman and Olga Khmelnitsky developed protocols for screening cowpea’s responses to salt and drought stress for soil-grown plants. Hayley Sussman screened a global natural diversity panel, consisting of 368 accessions, for their responses to drought, and is currently preparing the data for the Genome-Wide Association Study, to identify genetic components underlying drought stress tolerance.
Cowpea (Vigna unguiculata) is a legume crop native to West Africa, resilient to many biotic and abiotic factors. Hayley Sussman and Olga Khmelnitsky developed protocols for screening cowpea’s responses to salt and drought stress for soil-grown plants. Hayley Sussman screened a global natural diversity panel, consisting of 368 accessions, for their responses to drought, and is currently preparing the data for the Genome-Wide Association Study, to identify genetic components underlying drought stress tolerance.
Moreover, Hayley Sussman is also working on collecting material for DNA isolation and sequencing of the Nigerian cowpea diversity panel, which was shared with us by Dr Ramatu Aliyu (Ahmadu Bello University). By developing the Nigerian cowpea panel, in collaboration with Dr Aliyu, Dr Andrew Nelson and Dr Suzy Stickler (BTI), we aim to increase the number of identified SNPs, and develop pipelines for cowpea Genome-Wide Association Studies / RNA-sequencing, allowing quick trait evaluation and associations with genetic markers. This project, funded by the Triad foundation, will significantly contribute to future breeding efforts and accelerate the fundamental understanding of environmental resilience.
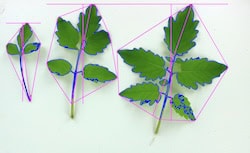
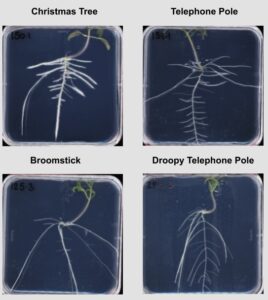 Solanum pimpinellifolium is the closest relative to cultivated tomato, and it is native to the west coast of South America. Different accessions of S. pimpinellifolium were collected from a wide variety of habitats and showed high diversity in their salt stress response in root architecture and salinity-induced changes therein. We recognized four visually distinct topologies of root architecture, that can be distinguished using Pareto Front optimality measures. We are working to expand the Pareto front calculations to improve the physiological representation of the constraints to root system architecture in the current model in collaboration with Dr Arjun Chandrasekhar (Southwestern University). Currently, Dr Maryam Rahmati-Ishka is investigating how we can alter distinct topologies using hormonal treatments, and what are the consequences of these topologies for sodium ion accumulation. Dr Rahmati-Ishka is also investigating the genetic components underlying salt stress sensitivity of lateral root development, by crossing two tomato lines with contrasting salt stress responses. Dr Julkowska is performing GWAS on collected root architecture data to identify candidate genes underlying root development and salt stress-induced changes. All the work in S. pimpinellifolium is currently focusing on salt stress. Hayley Sussman is responsible for the propagation of S. pimpinellifolium collection.
Solanum pimpinellifolium is the closest relative to cultivated tomato, and it is native to the west coast of South America. Different accessions of S. pimpinellifolium were collected from a wide variety of habitats and showed high diversity in their salt stress response in root architecture and salinity-induced changes therein. We recognized four visually distinct topologies of root architecture, that can be distinguished using Pareto Front optimality measures. We are working to expand the Pareto front calculations to improve the physiological representation of the constraints to root system architecture in the current model in collaboration with Dr Arjun Chandrasekhar (Southwestern University). Currently, Dr Maryam Rahmati-Ishka is investigating how we can alter distinct topologies using hormonal treatments, and what are the consequences of these topologies for sodium ion accumulation. Dr Rahmati-Ishka is also investigating the genetic components underlying salt stress sensitivity of lateral root development, by crossing two tomato lines with contrasting salt stress responses. Dr Julkowska is performing GWAS on collected root architecture data to identify candidate genes underlying root development and salt stress-induced changes. All the work in S. pimpinellifolium is currently focusing on salt stress. Hayley Sussman is responsible for the propagation of S. pimpinellifolium collection.
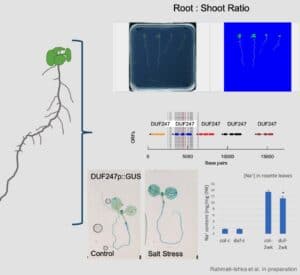 Salt stress rapidly impacts the growth and development of different plant organs to different extents, leading to alterations in plant development. Alterations to whole plant architecture, such as root-to-shoot ratio, have not been extensively explored thus far. Together with KAUST Visualization Lab, we developed a tool that quantifies green and white pixels, reflecting changes in root and shoot surface in Arabidopsis seedlings. Through forward genetics, we identified genetic components underlying salt stress-induced changes in root-to-shoot ratio including DUF247, encoding Domain of Unknown Function. Currently, Dr Rahmati-Ishka is examining the loss-of-function mutants of DUF247 using plate assays and soil-grown plants, as well as the subcellular localization of fluorophore-tagged DUF247 and GUS promoter fusion lines. Since the identified DUF247 is located in the genetic region containing other DUF247 genes, Hayley Sussman investigated the available loss-of-function mutants of the DUF247 genes located on Arabidopsis chromosome 3. Hayley Sussman generated transgenic lines targeting multiple DUF247 genes, as well as over-expression lines of specific DUF247 domains, that are currently being selected for homozygosity. Our preliminary results suggest that identified DUF247 is a negative regulator of Casparian strip formation and limits potential salt stress tolerance.
Salt stress rapidly impacts the growth and development of different plant organs to different extents, leading to alterations in plant development. Alterations to whole plant architecture, such as root-to-shoot ratio, have not been extensively explored thus far. Together with KAUST Visualization Lab, we developed a tool that quantifies green and white pixels, reflecting changes in root and shoot surface in Arabidopsis seedlings. Through forward genetics, we identified genetic components underlying salt stress-induced changes in root-to-shoot ratio including DUF247, encoding Domain of Unknown Function. Currently, Dr Rahmati-Ishka is examining the loss-of-function mutants of DUF247 using plate assays and soil-grown plants, as well as the subcellular localization of fluorophore-tagged DUF247 and GUS promoter fusion lines. Since the identified DUF247 is located in the genetic region containing other DUF247 genes, Hayley Sussman investigated the available loss-of-function mutants of the DUF247 genes located on Arabidopsis chromosome 3. Hayley Sussman generated transgenic lines targeting multiple DUF247 genes, as well as over-expression lines of specific DUF247 domains, that are currently being selected for homozygosity. Our preliminary results suggest that identified DUF247 is a negative regulator of Casparian strip formation and limits potential salt stress tolerance.
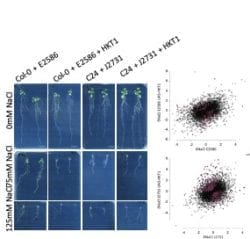 We discovered that high expression of Arabidopsis High-Affinity potassium (K+) Transporter (AtHKT1-1), sequestering sodium from the transpiration stream in root pericycle, results in increased salinity tolerance (Julkowska et al., 2017). Pericycle-specific over-expression of HKT1 resulted in reduced lateral root development under saline conditions. Interestingly, this effect seems to be specific to dicotyledonous species, as wheat lines with high HKT1 expression do not show a reduction in secondary lateral roots. Dr Julkowska examined the transcriptional changes underlying altered lateral root development downstream of HKT1 through comparative transcriptomic analysis of two lines with stelar over-expression of HKT1. Among the transcripts that were expressed differentially between UAS-HKT1 over-expression lines and their background lines, Dr Julkowska found genes involved in cell differentiation, ABA signalling and ion transport. Further validation of identified candidate genes using CRISPR-Cas9 knockouts is being currently being performed by Dr Julkowska, while over-expression lines of the genes are generated in Dr Mark Tester’s lab (KAUST). Further studies of generated mutant lines will enhance our understanding of processes controlling plant development under abiotic stress conditions and the importance of root architecture for salinity tolerance.
We discovered that high expression of Arabidopsis High-Affinity potassium (K+) Transporter (AtHKT1-1), sequestering sodium from the transpiration stream in root pericycle, results in increased salinity tolerance (Julkowska et al., 2017). Pericycle-specific over-expression of HKT1 resulted in reduced lateral root development under saline conditions. Interestingly, this effect seems to be specific to dicotyledonous species, as wheat lines with high HKT1 expression do not show a reduction in secondary lateral roots. Dr Julkowska examined the transcriptional changes underlying altered lateral root development downstream of HKT1 through comparative transcriptomic analysis of two lines with stelar over-expression of HKT1. Among the transcripts that were expressed differentially between UAS-HKT1 over-expression lines and their background lines, Dr Julkowska found genes involved in cell differentiation, ABA signalling and ion transport. Further validation of identified candidate genes using CRISPR-Cas9 knockouts is being currently being performed by Dr Julkowska, while over-expression lines of the genes are generated in Dr Mark Tester’s lab (KAUST). Further studies of generated mutant lines will enhance our understanding of processes controlling plant development under abiotic stress conditions and the importance of root architecture for salinity tolerance.
 Studying the drought resilience using a non-destructive experimental setup poses significant opportunities. Traditionally, soil-grown plants are evaluated based solely on the above-ground portions due to easier accessibility. Aeroponics technology stands out as a game-changer, as it provides easy access to roots and shoots for imaging, and offers precise control over water supply. In collaboration with Prof. Perrinne Periot (CU Enngineering) we are developing aeroponics system where roots are misted with a nutrient solution. We have successfully mimicked drought conditions, eliciting physiological responses similar to those of drought-stressed plants grown in soil. As aeroponics allows for rapid changes in water availability, it will permit differentiation between water conservation and true resilience strategies within tolerant genotypes.
Studying the drought resilience using a non-destructive experimental setup poses significant opportunities. Traditionally, soil-grown plants are evaluated based solely on the above-ground portions due to easier accessibility. Aeroponics technology stands out as a game-changer, as it provides easy access to roots and shoots for imaging, and offers precise control over water supply. In collaboration with Prof. Perrinne Periot (CU Enngineering) we are developing aeroponics system where roots are misted with a nutrient solution. We have successfully mimicked drought conditions, eliciting physiological responses similar to those of drought-stressed plants grown in soil. As aeroponics allows for rapid changes in water availability, it will permit differentiation between water conservation and true resilience strategies within tolerant genotypes.
 Understanding metabolic contributions to drought stress resilience in cowpea and other legume species.
Understanding metabolic contributions to drought stress resilience in cowpea and other legume species.
Lab Members

Molly Christel

Emily Miller

Ruth Nichols

Aaron Quan

Sofia Trujillo Ortigoza
Research Technician II

Akhilesh Sharma
PhD student
Past Lab Members

Aminu Aliyu

Aparna Srinivasan

Hayley Sussman

Olga Khmelnitsky

Leanna Skeete

Jade Milton
In the News

BTI, Meiogenix, and FFAR Announce $2 Million Breakthrough Tomato Genetics Collaboration
Research Lays the Foundation for Breakthroughs in Global Food Security In a landmark $2 million initiative, the Boyce Thompson Institute (BTI) and biotechnology company Meiogenix have launched a collaboration to develop drought- and disease-resistant tomatoes by tapping...
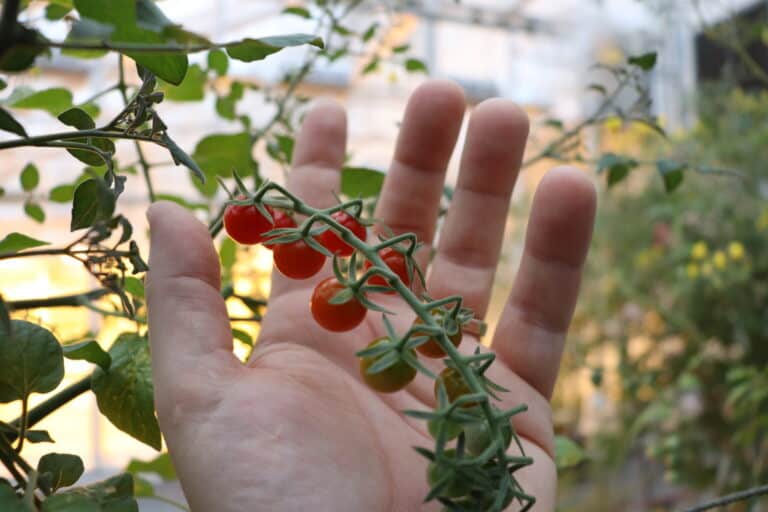

As weather patterns shift and soil salinity increases in many agricultural areas, finding crops that can thrive in these challenging conditions is crucial. Cultivated tomatoes, while delicious, often struggle in...
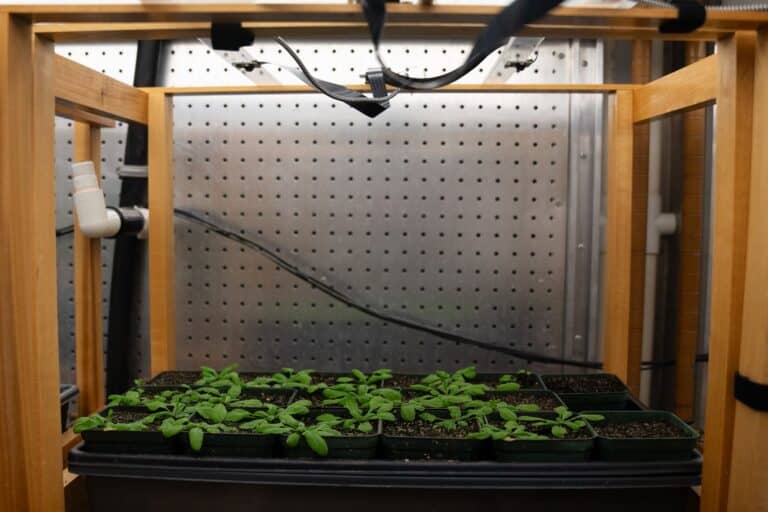

Phenotyping, which involves assessing observable plant characteristics, is crucial for understanding plant development and response to environmental stresses. Traditional methods are often cumbersome, costly, and destructive, limiting research scope and...
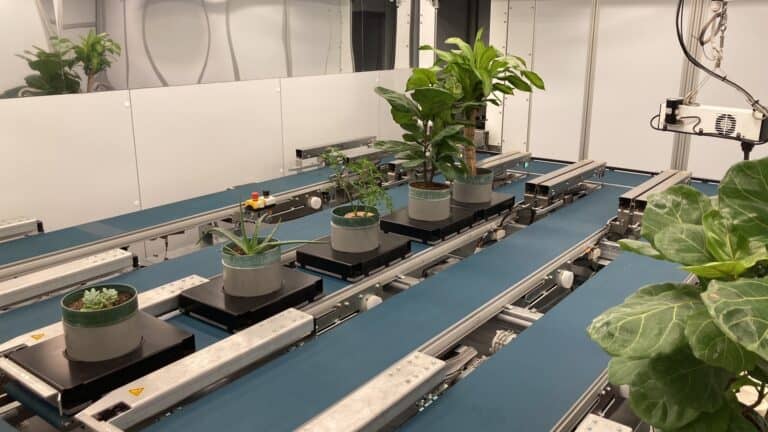

BTI is proud to announce the opening of its cutting-edge plant phenotyping facility, PhenoSight. This new facility holds significant potential for delivering unparalleled insights into plant growth, development, and response...

Cluster Hire Yields Three New Faculty Members
Boyce Thompson Institute is pleased to announce the hiring of three faculty members as part of its new and innovative “cluster hire” approach. Out of 113 applicants, the three people...
Internships
BTI offers a summer research experience program for undergraduate and high school students.
Previous Interns

Katie Carson
The role of genes encoding pericycle-specific expression in development of lateral roots and salinity tolerance
Salinity stress is a contributing factor to lowering crop yield due to the accumulation of sodium ions in the shoot and interfering with photosynthesis as well as a range of other independent processes. During the early stages of plant development, lateral roots develop from the pericycle, the cell layer enclosing the vascular tissue. The pericycle is important for processes regulating water and ion transport. Previously, we observed that Arabidopsis high-affinity potassium transporter (HKT1) contributes to salt tolerance and lateral root development. Here, we hypothesize that other genes with pericycle-specific expression are key to lateral root development under salt conditions. Apart from Arabidopsis’ one HKT1 gene, two HKT1 genes (HKT1;1 and HKT1;2) have been identified in tomato, however, their tissue-specific expression has yet to be determined. In this study, we aimed to determine the tissue-specific expression of HKT1;1 and HKT1;2 through cloning their respective promoters into GreenGate expression vectors. In the future, these constructs will be transformed into Agrobacterium rhizogenes and ultimately induce hairy root transformations in tomatoes to localize the pericycle-specific expression of HKT1;2. Additionally, we screened 47 Arabidopsis mutants targeting genes with pericycle-specific expression. The mutants were evaluated for root architecture with and without salt stress. 13 mutants with altered salt stress resilience underwent further screening in PhenoSite Facility at BTI. The results collected within this study will increase our understanding of genes that are specifically expressed in the pericycle, and their contribution to salt stress resilience. This and future studies will further contribute to a more targeted approach to cultivating crops with low-quality water and in areas with no or limited access to freshwater.
I am extremely grateful for the opportunity to work in Magda Julkowska’s lab at BTI and Cornell University this summer. I came to Ithaca lacking a lot of microbiology laboratory experience and I was excited to be out of my comfort zone and to learn everything I could. Over the past ten weeks, my discomfort in new challenges such as cloning via the GreenGate method and the use of R for data analysis has pushed me to become a better biologist. Magda went to great lengths to ensure my experience was the best it could be and it shines in the bittersweet sentiment I am feeling as my time here comes to an end. She has been an amazing mentor; she encouraged me to learn while supporting me and making it okay to make mistakes. My time here has nurtured and simultaneously tested my passion for plant sciences, and I am emerging from it more confident than ever that plant biology is the field I aim to persue.
Intern Info

Annie Voss
Molecular Mechanics Underlying Tepary Bean Responses to Drought Stress
As climate change progresses, rising global temperatures continue to increase stress on the world’s agricultural systems. The common bean (whose varieties include green, navy, black, kidney, and pinto beans) has proven particularly susceptible to these changes. In fact, over two-thirds of the world’s legumes are expected to be at risk (Zaulda et al., 2022). The tepary bean, a sister species of the common bean, is indigenous to the southwestern United States and Mexico and has emerged as a viable source of genetic variation for increasing the common bean’s resistance to abiotic stress. Previously, we have identified genes that underlie tepary bean’s resistance to drought stress using forward genetics. Here, we conducted experiments to validate individual genes’ contributions towards drought resilience. Firstly, we successfully validated a protocol for cowpea mosaic virus-induced gene silencing (VIGS) in tepary bean and cowpea. Secondly, we evaluated tepary bean accessions previously classified as stress-resilient or susceptible in the PhenoSight facility for their responses to drought stress. Finally, we screened T-DNA insertion mutants in Arabidopsis of genes homologous to tepary bean genes associated with drought stress resilience. Our results were promising on several fronts. Thanks to our successful validation of the VIGS protocol and evaluation of tepary accessions as tolerant and susceptible to drought stress, we are now targeting drought stress genes using VIGS in tepary beans. Further, our evaluation of Arabidopsis mutants revealed the potential role of several genes in response to osmotic stress at a large evolutionary scale, which may allow us to gain further insight into the broad application of our findings. These and future studies will help us advance our understanding of what makes tepary bean so resilient to drought stress, allowing the identification of future breeding targets in legumes and beyond, and contributing to greater global food security.
I am immensely grateful to have been placed in the Julkowska Lab. The culture of Magda’s lab and the effort she put into her mentorship made me feel welcomed, respected, and safe to make mistakes or not know it all right away. When I arrived, I wrote down three things I hoped to accomplish during this experience: a better understanding of my career goals, to gain confidence, and to increase my knowledge in depth, breadth, and application. I know I have accomplished all three of these goals. This REU has helped me learn about careers in plant science and agriculture I was previously unaware of. The daunting goal of exploring grad school seems somewhat lessened by the information and experience I gained, and I feel much more at home in the lab setting. While no experience is without its challenges, Magda and the BTI staff’s tireless efforts have made the experience worthwhile, and I am truly thankful.
Intern Info

Ting Fei
“Exploring salt stress on Arabidopsis thaliana growth with DUF247 genes”
Project Summary:
Salt stress is detrimental to plant growth. Climate change, floods, irrigation, runoff, and silting all introduce increased amounts of salt into the soil that pose a serious threat to the survival of plants. With our growing population, along with unstable food security, the need for improvements in agriculture is increasing. In a previous study, it was observed that the natural variation in salt stress-induced changes in root-to-shoot ratio corresponds to a locus-encoding domain of unknown function gene, AT3G50160, within the DUF247 family. However, the identified DUF247 locus contains other DUF247 genes that share high sequence similarity to our prime candidate, AT3G50160. Our objective is to determine the physiological relevance of the AT3G50160 gene, as well as its neighboring genes, in response to salt stress by observing the root system architecture on plates and rosette size in soil. By understanding which genes contribute to salt stress tolerance, we can engineer plants that are more resilient to high concentrations of salt.
My Experience:
Throughout my summer at BTI, I was able to better understand what it’s like to work in a research environment and what I would like to do in the future. In such a supportive and friendly lab, with Hayley Sussman as my mentor, I was able to grow with their expertise while also feeling comfortable with making mistakes along the way. I got to improve on relevant skills, such as communication and time management, as well as learn how to use a lot of new research tools, such as ImageJ and R. This internship has allowed me to greatly expand my knowledge and I am looking forward to applying what I learned later on. Having this opportunity to further explore the world of science is an experience I will always be grateful for.
Intern Info

Yalmarie Numan Vazquez
“Hormonal modulation of root system architecture in wild tomatoes under salt stress”
Project Summary:
The increasing human population and reduction in land available for cultivation are threats to agricultural sustainability. This causes environmental stress like soil salinity which alters plant architecture and growth rate, affecting overall plant productivity. Thus, it is imperative to develop more salt-tolerant plants for sustainable agriculture. Since roots are at the interface of saline soil, many root-related architectural traits have been the target of breeding programs for improving crop tolerance and yield. Solanum pimpinellifolium (wild tomato) is the closest relative to Solanum lycopersicum (cultivated tomato) and is an important source for genetic improvement since it goes through adaptation to tolerate any abiotic stress. Previously, Julkowska et al. visually classified four distinct root topologies in terms of the salt stress response by screening root system architecture of 230 S. pimpinellifolium accessions. However, it still remains to be determined 1) how hormonal signaling is involved in creating distinct root topologies, and 2) what is the biological significance of these root topologies in the overall salt stress response. In our experiment, we compared the effect of two classical plant hormones, abscisic acid and ethylene, on the root system architecture of two wild tomato accessions and one cultivated tomato accession. For studying root system architecture, the seedlings are germinated in a control plate for five days and then transferred into treatment plates. Then, the plates start to be scanned for five consecutive days. The images obtained are analyzed through the plugin SmartRoot in the program ImageJ for root tracing, followed by root architectural traits analysis in R.
My Experience:
This experience made me grow both professionally and personally. Coming from such a small island like Puerto Rico and being so far away from home for a whole summer was a real challenge but it was definitely worth it. I made unforgettable friends and memories that forever will live with me. Being an intern at Boyce Thompson Institute certainly changed my vision of what plant biology research is like. It allowed me to enhance my confidence in a research setting, sharpen my critical thinking, learn about new research tools, and develop new skills that will be of high value for my future. In addition to this, I had the opportunity to meet and work with amazing people in my research lab that helped me identify my research interests and define my career goals. This internship provided me with the support and mentorship necessary to approach graduate school applications on the right foot. Overall, I learned valuable lessons and had a fruitful experience that will carry with me.
Intern Info

Zoe James
“Investigation of the role of DUF247 gene in the regulation of salt stress in Arabidopsis Thaliana“
Project Summary:
Salt stress alters plant architecture by impacting the growth and development of different organs, such as main and lateral root growth. Other alterations in plant architecture, such as changes in root-to-shoot ratio are hypothesized to contribute to salt tolerance. Previous screening into salt stress-induced changes in root-to-shoot ratio identified underlying genetic components using Genome Wide Association Study. One of the identified loci was DUF247, encoding a protein with Domain of Unknown Function. The role of DUF247 in regulating root to shoot ratio, as well as general salt tolerance in Arabidopsis is unknown. We aimed to characterize the DUF247 loss-of-function mutant to identify the potential link between this gene and its contribution to the regulation of salt stress response. We compared the duf247 mutant and wild-type (Col-0) plants with or without salt treatment for seed germination and seedling survival rates. Further, we monitored dynamic changes in growth in the above-ground tissues for the soil grown plants for both duf247 mutant and Col-0 plants, and measured the evapotranspiration rate, growth rate as well as rosette fresh weight. Overall, our results showed no significant difference between Col-0 and duf247 mutant plants in terms of germination and survival assays. The future plan will be to repeat the plate experiments using higher concentrations of salt.
My Experience:
My summer internship in the Julkowska lab at BTI has been a fulfilling and extremely valuable experience. My mentor, Maryam Rahmati Ishka, always positively supported me and allowed me to build independence in the lab with confidence in my work. Being welcomed into lab meetings, weekly seminars, and journal clubs provided me with a holistic experience as a plant scientist beyond working at the bench. I learned techniques such as RNA extraction, Real-time qPCR, cDNA synthesis along with other protocols and methods for my experiments. I also enjoyed organizing my results and data onto my poster and preparing to present it. Through the many opportunities to learn about future studies and careers in science, I feel more confident in my future plans for college and am excited to conduct research in the future.
Intern Info






