Inspiration
Kewal Shah, our teammate in the MS Human-Computer Interaction program, spent time surveying physical therapy clinics for areas where tools and data science could improve the patient experience. He found that patients and loved ones would consistently ask for reports and progress updates, which required the physical therapists to re-review notes and have to translate medical jargon. Rachel Calder, a MS student in Bioinformatics was curious about how pervasive this issue was within the healthcare field, and discovered a need for subsects of the medical field where physician efficiency is crucial (Kannampallil et al 2018).
The team was inspired to create a tool to reduce the time healthcare professionals (HPs) spend on electronic health records (EHR). The project used Kewal's front-end skills, Rachel's health data science skills, and the back-end skills of Computer Science undergraduate students Hailey Ho and Gyandeep Reddy Vulupala. The project was named CareA11y to combine the concepts of improving the efficiency of patient's understanding of care while following standard accessibility (a11y) practices during development. We verified that the text was easy to read and accessible by checking Contrast using the WebAIM tool and using appropriate aria attributes to support users with disabilities.
What it does
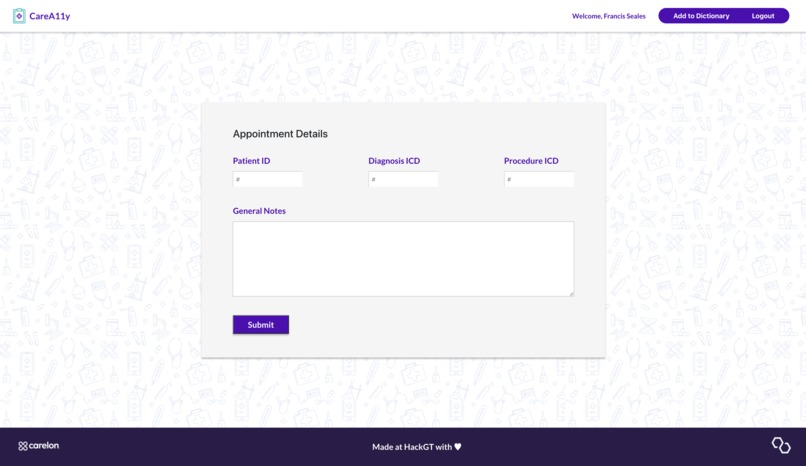
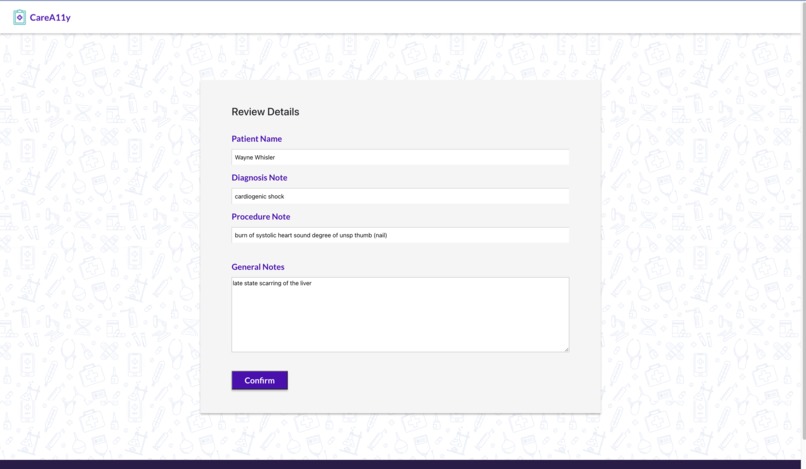
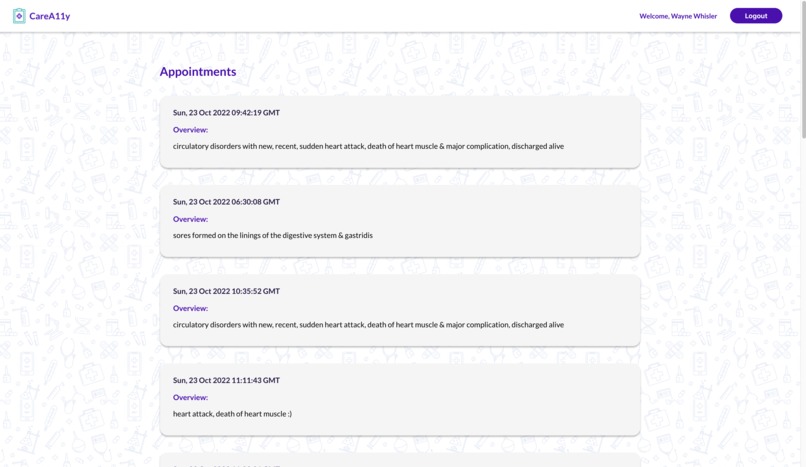
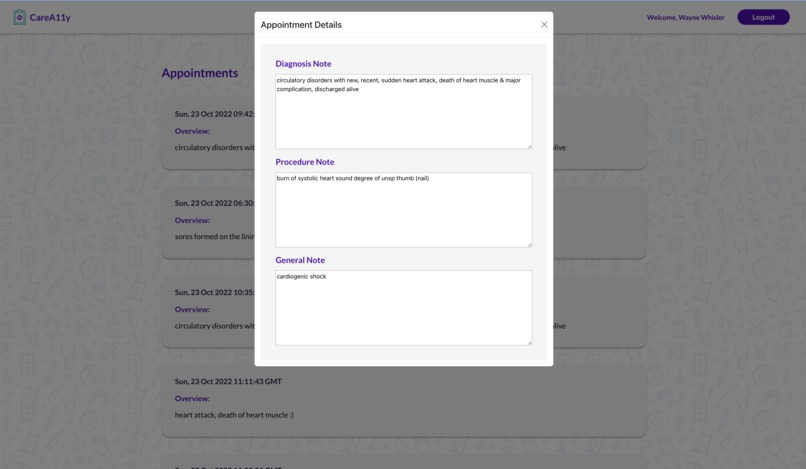
CareA11y allows HPs to input notes with medical jargon (often shorthand) for a particular patient. They can also input ICD diagnosis and procedure codes for further translation which are used to query live websites for associated text. All medical text is sent to the Google Cloud API identifies medical jargon terms. These terms are then translated via an aggregated specialized text translator library. For any given doctor visits, there are 3 different translated notes - general/summary visit notes, procedures, and diagnoses. The HPs can review the text for accuracy before approving it for the patient to view. To enrich our translation dataset, the HPs can also add in their own technical-to-layman value pairs. A patient can log in, select a desired appointment, and review the associated notes.
How we built it
We designed the high-fidelity design prototype using Figma and exported the designs to Zeplin to get the design specifications. We built the frontend using React, The backend website was built using Python (Flask), PostgreSQL, and Heroku. The database was supported by PostgreSQL, the APIs were developed by leveraging the Flask library in Python. We extracted medical terms from the text using Google Cloud Healthcare Natural Language API and replaced those texts with layman's terms using a compiled dataset.
Challenges we ran into
Data Acquisition: The first challenge was aggregating meaningful EHRs given HIPAA (Health Insurance Portability and Accountability Act of 1996) restrictions. We decided to use the MIMIC-III open source database information, but noticed text descriptions of patient encounters could be only a few words long. Therefore, we added additional data based on ICD code descriptions.
Backend Development: Team members had experience with backend development on local machines during hackathons, however, it was a challenge to deploy the code to a cloud platform. We chose to host our code online to make it easier and more flexible to utilize CRUD and other processing APIs in the front end service. We ran into numerous issues:
- We had to explore and configure API clients for both AWS Comprehend Medicine API and Google Cloud Healthcare Natural Language API to evaluate their accuracy.
- The deployment required the use of git where two remote repositories were managed at the same time (i.e., GitHub and the cloud platform).
- The git references were hard to navigate.
- There were instances where the code was performing as expected on the local machine, but crashed on the cloud platform.
Debugging the root cause of issues was challenging, but it was also a great way to learn from our mistakes and gain experience.
Accomplishments that we're proud of
We are proud of creating a fully functional frontend and backend product with many avenues to expand upon. We are also proud of being able to work together for 36 hours, having just met. Our skills complemented each other.
What we learned
We learned that we are capable of going more than 24 hours without sleep. One a serious note, each participant pushed their skill sets and would alternate teaching and learning different tools and languages from each other to efficiently break down the work.
What's next for CareA11y
CareA11y is interested in working with larger-text and more complex EHRs, and implementing machine learning and natural language processing techniques to improve readability scores (like Flesch) and allow for model re-training based on HPs feedback edits to continue to minimize the time spent away from patient care. For example, if a HP approves a note, that note would be stored as a 'good' note for that diagnosis.
References
Johnson, A. E. W., Pollard, T. J., Shen, L., Lehman, L. H., Feng, M., Ghassemi, M., Moody, B., Szolovits, P., Celi, L. A., & Mark, R. G. (2016). MIMIC-III, a freely accessible critical care database. Scientific data, 3, 160035.
Kannampallil, T., Denton, C., Shapiro, J., & Patel, V. (2018). Efficiency of emergency physicians: Insights from an observational study using EHR log files. Applied Clinical Informatics, 09(01), 099–104. https://doi.org/10.1055/s-0037-1621705

Log in or sign up for Devpost to join the conversation.