GermAlert: Predicting Outbreaks Before They Happen
What Inspired Me
I am a 4th semester Bioinformatics student from Pakistan — a country where disease outbreaks are not abstractions in a textbook. They are news alerts, community crises, and preventable deaths. Growing up, I watched how cholera, dengue, and typhoid repeatedly overwhelmed hospitals that had no warning they were coming.
The question that haunted me was simple: why do we always respond after the damage is done?
When I discovered that wastewater-based epidemiology (WBE) had successfully detected COVID-19 surges in cities like Amsterdam and New York — days before hospitals saw a single new case — I realized the technology existed. What did not exist was access to it for the countries that needed it most.
That gap became GermAlert.
What I Learned
Building GermAlert taught me that public health is a data problem as much as a medical one. The genomic sequences sitting in NCBI's Sequence Read Archive, the climate anomalies streaming from NASA's POWER API, and the outbreak records archived in WHO GLASS — all of this intelligence exists. It is just disconnected, inaccessible, and untranslated for the people who need it.
I learned how resistance genes like blaKPC, blaNDM, and mcr-1 move through wastewater before clinical cases appear. I learned that a spike in rainfall combined with elevated microbial load is not a coincidence — it is a signal. And I learned that machine learning does not need to be complicated to be powerful.
The core prediction model uses a Random Forest classifier, which works on a beautifully simple principle: aggregate the decisions of many weak learners into one strong prediction. Mathematically, for $T$ trees, the final prediction is:
$$\hat{y} = \text{mode}\left({h_t(x)}_{t=1}^{T}\right)$$
where $h_t(x)$ is the prediction of the $t$-th decision tree for input feature vector $x$.
The feature vector for each region at time $t$ is:
$$x_t = \left[\text{PathogenLoad}_t,\ \Delta T_t,\ R_t,\ F_t,\ G_t\right]$$
where $\Delta T_t$ is the temperature anomaly, $R_t$ is the rainfall index, $F_t$ is the flood risk score, and $G_t$ is the genomic resistance signal strength — all normalized to $[0, 1]$.
The model outputs a risk score $s \in [0, 100]$, calibrated as:
$$s = 100 \times P\left(\text{outbreak within 21 days} \mid x_t\right)$$
This was the most intellectually exciting part of the build — taking biology, atmospheric science, and machine learning and collapsing them into a single number that a health minister can act on.
How I Built It
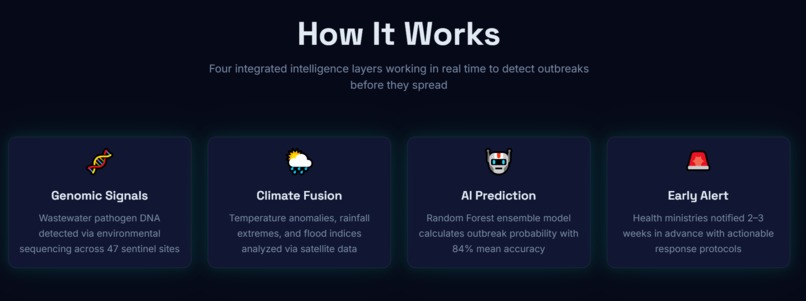
GermAlert was built in four layers:
Layer 1 — Data Pipeline
Using Python and pandas, I built automated scripts to pull
wastewater pathogen genomic sequences from NCBI SRA and WHO
GLASS, and real-time climate variables from the NASA POWER API.
These were merged into a unified regional dataset indexed by country
and timestamp.
Layer 2 — AI Prediction Engine
The merged dataset was fed into a scikit-learn Random Forest classifier trained on 15 years of WHO historical outbreak records. The model was validated using stratified k-fold cross-validation:
$$\text{Accuracy} = \frac{1}{k}\sum_{i=1}^{k} \frac{TP_i + TN_i}{TP_i + TN_i + FP_i + FN_i}$$
The result was a regional outbreak risk score updated every 6 hours as new data streams arrived.
Layer 3 — Dashboard
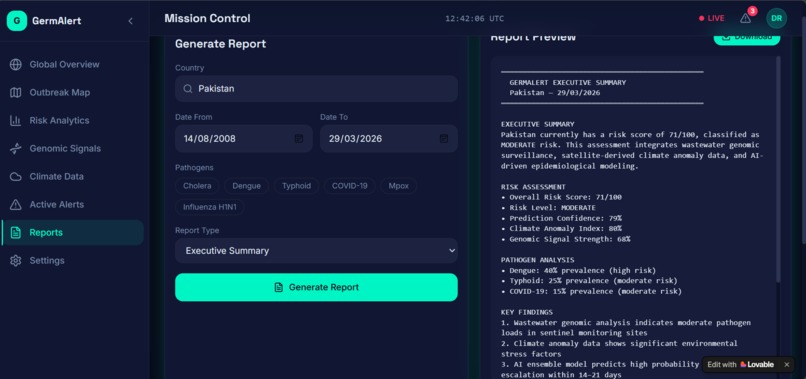
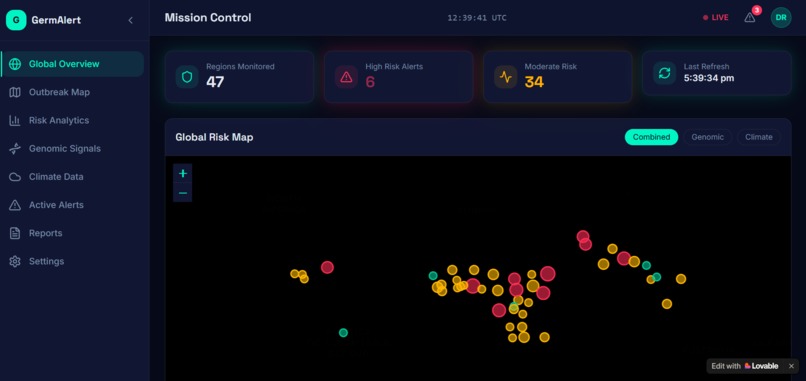
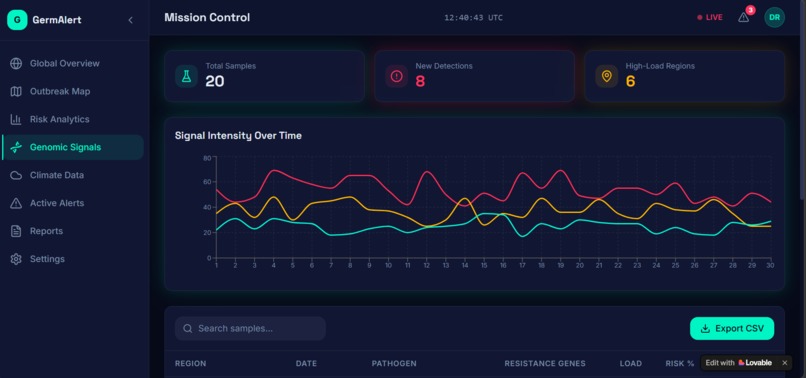
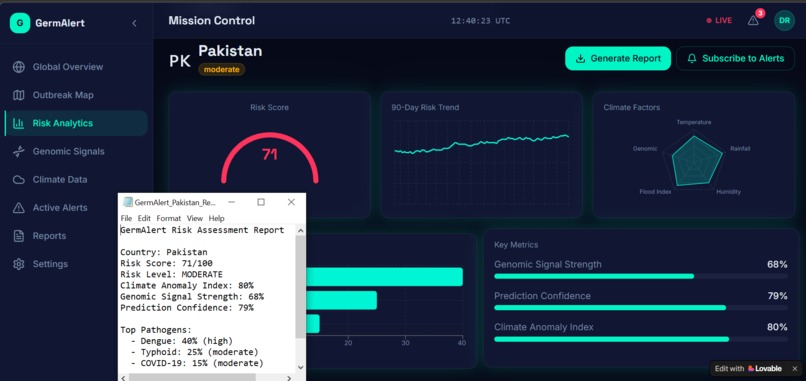
The prediction outputs were visualized using Streamlit as an interactive dashboard — showing a world map with color-coded risk zones, 90-day trend charts, pathogen breakdowns, and country-level drill-downs.
Layer 4 — Automation and Delivery
n8n was used to automate the entire pipeline — scheduling daily data pulls, triggering model retraining when new samples arrived, and dispatching email alerts to health ministries when risk scores crossed critical thresholds. Lovable was used to wrap the dashboard in a professional, accessible frontend designed for non-technical health workers.
Challenges I Faced
1. I Was a Beginner
I will not pretend otherwise. At the start of this project, I had basic Python knowledge and no experience building end-to-end AI systems. Every component — API integration, machine learning pipelines, dashboard deployment — was new territory. The challenge was not just technical. It was psychological: learning to be comfortable with not knowing, and trusting that understanding would come through building.
2. Data Scarcity for Low-Income Countries
The cruel irony of this project is that the countries with the highest outbreak burden have the least surveillance data. NCBI SRA has rich wastewater genomic records for Europe and North America — but Sub-Saharan Africa and South Asia are dramatically underrepresented. This forced me to think carefully about model generalizability and the ethics of predicting risk in regions where data gaps themselves are a form of inequity.
3. Fusing Heterogeneous Data Sources
Genomic signals and climate variables exist in completely different formats, scales, and temporal resolutions. Aligning a DNA sequence confidence score with a rainfall anomaly index required careful normalization and feature engineering. Getting the model to treat both signal types as equally informative — rather than overfitting to whichever had higher variance — was a real technical challenge.
4. Making It Usable for Non-Experts
The most important user of GermAlert is not a data scientist. It is a district health officer in Nigeria who has never opened a terminal window. Every design decision — from the color-coded risk scores to the one-click PDF report — had to be made with that person in mind. Translating machine learning confidence intervals into plain language that drives real decisions is harder than building the model itself.
What GermAlert Means to Me
This is my first real project. Not a class assignment, not a tutorial — a genuine attempt to build something that could matter.
The $1.27$ million people who die from antimicrobial resistance every year are not a statistic to me. They are people in communities like mine, in countries like mine, failed by systems that could have warned us earlier.
GermAlert is my answer to that failure. It is imperfect, it is early, and it has a long way to go. But it is real — and that is where everything begins.
"The best time to predict an outbreak is before it starts. The second best time is now."
Built With
- and-alert-dispatch-design-and-frontend-lovable-for-ui-and-ux
- card-database-automation-n8n-for-automated-data-pipeline
- chart.js
- css
- framer-motion
- git
- github-cloud-and-apis-nasa-power-api
- html
- javascript-frameworks-and-libraries-scikit-learn
- languages-python
- leaflet.js
- leaflet.js-for-interactive-maps
- n8n
- ncbi-sra-api
- numpy
- our-world-in-data-ai-and-machine-learning-random-forest-classifier
- pandas
- react-platforms-and-tools-lovable
- scheduled-pulls
- scikit-learn-pipeline
- stratified-k-fold-cross-validation-data-and-databases-ncbi-sequence-read-archive
- streamlit
- vs-code
- who-glass
- who-glass-database

Log in or sign up for Devpost to join the conversation.